library(ggplot2)
ggplot(cdc) +
aes(year, cases) +
geom_point() +
geom_line()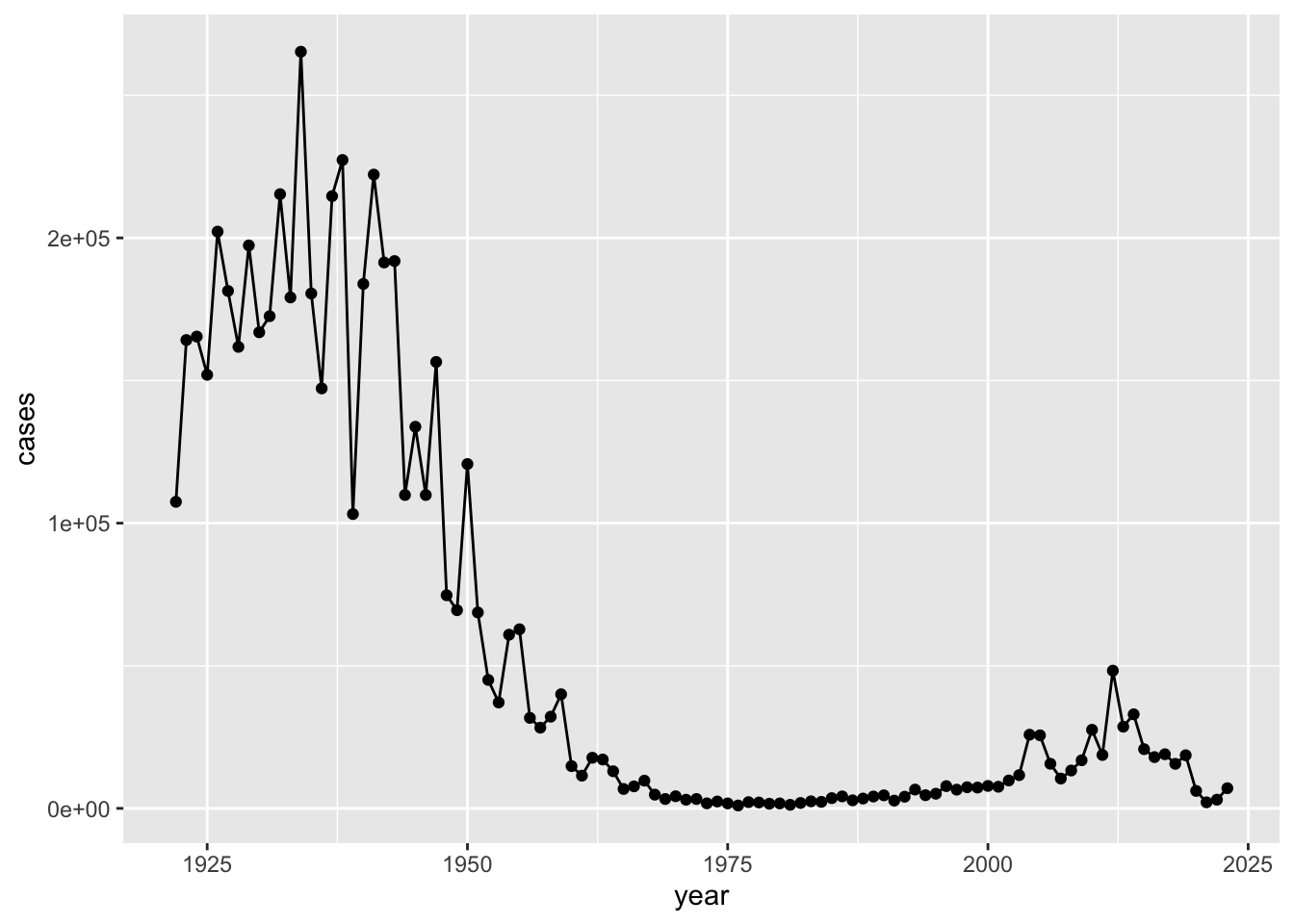
Pertussis (a.k.a whooping cough) is a common lung infection caused by the bacteria B. Pertussis.
This can infect all ages but is most severe for those under 1 year of age.
The CDC track the number of reported cases in the US
We can “scrape” this data with the datapasta package.
Q1. Make a plot of
yearvscases
library(ggplot2)
ggplot(cdc) +
aes(year, cases) +
geom_point() +
geom_line()
Q2. Add some major milestones including the first wP vaccine rollout (1946), the switch to the newer aP vaccine (1996).
ggplot(cdc) +
aes(year, cases) +
geom_point() +
geom_line() +
geom_vline(xintercept = 1946, col = "blue", lty = 2) +
geom_vline(xintercept = 1996, col = "red", lty = 2) +
geom_vline(xintercept = 2020, col = "darkgreen", lty = 2) 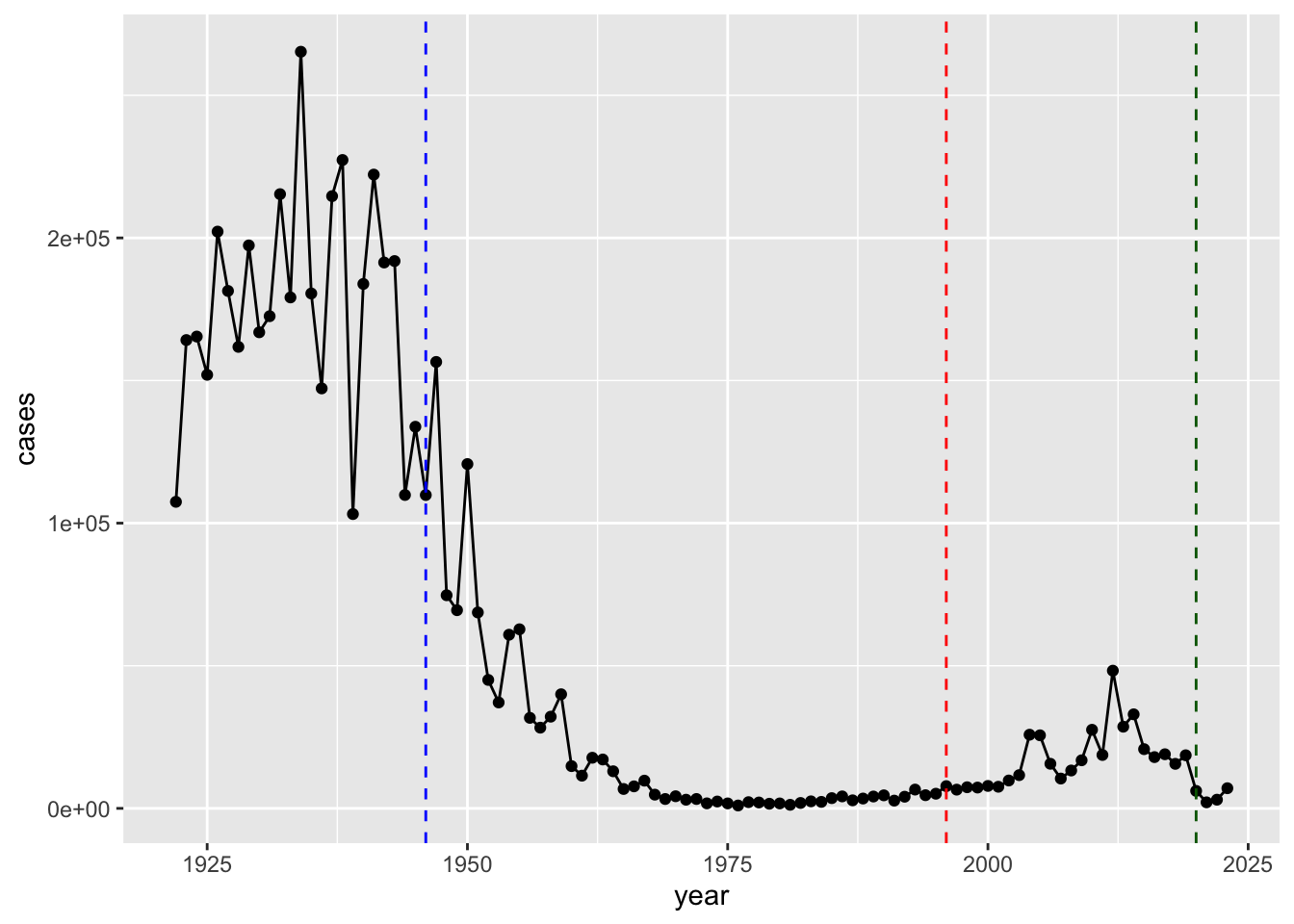
Q3. Describe what happened after the introduction of the aP vaccine? Do you have a possible explanation for the observed trend?
There were high case numbers in the pre 1940s, then came the introduction of the aP vaccine. This dramatically reduced the number of cases. Differing from the wP vaccine, the aP vaccine needs a booster vaccine after around 10 years. Around the 2000s we can see a rise in cases, most likely due to people not getting booster vaccines. Around 2020, we can see cases drop due to quarantine for COVID-19, but rise again likely due to anti-vaxxers from the fear of the COVID-19 vaccine. The aP induced protection wanes faster than wP.
Why is this vaccine-preventable disease on the upswing? To answer this question we need to investigate the mechanisms underlying waning protection against pertussis. This requires evaluation of pertussis-specific immune responses over time in wP and aP vaccinated individuals.
Computational Models of Immunity - Pertussis Boost project aims to provide the scientific comunity with this very information.
They make their data available via JSON format APIs. We can read this in R with the read_json() function from the jsonlite package:
library(jsonlite)
library(ggplot2)
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(lubridate)
Attaching package: 'lubridate'The following objects are masked from 'package:base':
date, intersect, setdiff, unionsubject <- read_json("https://www.cmi-pb.org/api/subject", simplifyVector = TRUE)
specimen <- read_json("https://www.cmi-pb.org/api/v5_1/specimen",
simplifyVector = TRUE)
ab_titer <- read_json("https://www.cmi-pb.org/api/v5_1/plasma_ab_titer",
simplifyVector = TRUE)
meta <- inner_join(specimen, subject, by = "subject_id")
abdata <- inner_join(ab_titer, meta, by = "specimen_id")Q4. How many aP and wP infancy vaccinated subjects are in the dataset?
table(subject$infancy_vac)
aP wP
87 85 There are 87 aP and 85 wP infancy vaccinated subjects in this dataset.
Q5. How many Male and Female subjects/patients are in the dataset?
table(subject$biological_sex)
Female Male
112 60 There are 112 female and 60 male subjects/patients in this dataset.
In terms of race and gender is this dataset representative of the US population?
Q6. What is the breakdown of race and biological sex (e.g. number of Asian females, White males etc…)?
table(subject$race, subject$biological_sex)
Female Male
American Indian/Alaska Native 0 1
Asian 32 12
Black or African American 2 3
More Than One Race 15 4
Native Hawaiian or Other Pacific Islander 1 1
Unknown or Not Reported 14 7
White 48 32head(specimen) specimen_id subject_id actual_day_relative_to_boost
1 1 1 -3
2 2 1 1
3 3 1 3
4 4 1 7
5 5 1 11
6 6 1 32
planned_day_relative_to_boost specimen_type visit
1 0 Blood 1
2 1 Blood 2
3 3 Blood 3
4 7 Blood 4
5 14 Blood 5
6 30 Blood 6head(ab_titer) specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgE FALSE Total 1110.21154 2.493425
2 1 IgE FALSE Total 2708.91616 2.493425
3 1 IgG TRUE PT 68.56614 3.736992
4 1 IgG TRUE PRN 332.12718 2.602350
5 1 IgG TRUE FHA 1887.12263 34.050956
6 1 IgE TRUE ACT 0.10000 1.000000
unit lower_limit_of_detection
1 UG/ML 2.096133
2 IU/ML 29.170000
3 IU/ML 0.530000
4 IU/ML 6.205949
5 IU/ML 4.679535
6 IU/ML 2.816431library(lubridate)What is today’s date (at the time I am writing this obviously)
today()[1] "2026-03-16"How many days have passed since new year 2000
today() - ymd("2000-01-01")Time difference of 9571 daysWhat is this in years?
round(time_length(today() - ymd("2000-01-01"), "years"), 2)[1] 26.2To analyze this data we need to first “join” the different tables so we have all the data in one place, not spread across different tables.
We can use the *_join() family of functions from dplyr to do this.
library(dplyr)
meta <- inner_join(subject, specimen)Joining with `by = join_by(subject_id)`head(meta) subject_id infancy_vac biological_sex ethnicity race
1 1 wP Female Not Hispanic or Latino White
2 1 wP Female Not Hispanic or Latino White
3 1 wP Female Not Hispanic or Latino White
4 1 wP Female Not Hispanic or Latino White
5 1 wP Female Not Hispanic or Latino White
6 1 wP Female Not Hispanic or Latino White
year_of_birth date_of_boost dataset specimen_id
1 1986-01-01 2016-09-12 2020_dataset 1
2 1986-01-01 2016-09-12 2020_dataset 2
3 1986-01-01 2016-09-12 2020_dataset 3
4 1986-01-01 2016-09-12 2020_dataset 4
5 1986-01-01 2016-09-12 2020_dataset 5
6 1986-01-01 2016-09-12 2020_dataset 6
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 1 1 Blood
3 3 3 Blood
4 7 7 Blood
5 11 14 Blood
6 32 30 Blood
visit
1 1
2 2
3 3
4 4
5 5
6 6abdata <- inner_join(ab_titer, meta)Joining with `by = join_by(specimen_id)`head(abdata) specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgE FALSE Total 1110.21154 2.493425
2 1 IgE FALSE Total 2708.91616 2.493425
3 1 IgG TRUE PT 68.56614 3.736992
4 1 IgG TRUE PRN 332.12718 2.602350
5 1 IgG TRUE FHA 1887.12263 34.050956
6 1 IgE TRUE ACT 0.10000 1.000000
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 UG/ML 2.096133 1 wP Female
2 IU/ML 29.170000 1 wP Female
3 IU/ML 0.530000 1 wP Female
4 IU/ML 6.205949 1 wP Female
5 IU/ML 4.679535 1 wP Female
6 IU/ML 2.816431 1 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
5 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
6 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1Q. What antibody isotypes are measured for these patients?
table(abdata$isotype)
IgE IgG IgG1 IgG2 IgG3 IgG4
6698 7265 11993 12000 12000 12000 Q. What antigens are reported?
table(abdata$antigen)
ACT BETV1 DT FELD1 FHA FIM2/3 LOLP1 LOS Measles OVA
1970 1970 6318 1970 6712 6318 1970 1970 1970 6318
PD1 PRN PT PTM Total TT
1970 6712 6712 1970 788 6318 Let’s focus on the IgG isotype and make a plot of MFI_normalized for all antigens.
igg <- abdata |>
filter(isotype == "IgG")
head(igg) specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgG TRUE PT 68.56614 3.736992
2 1 IgG TRUE PRN 332.12718 2.602350
3 1 IgG TRUE FHA 1887.12263 34.050956
4 19 IgG TRUE PT 20.11607 1.096366
5 19 IgG TRUE PRN 976.67419 7.652635
6 19 IgG TRUE FHA 60.76626 1.096457
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 IU/ML 0.530000 1 wP Female
2 IU/ML 6.205949 1 wP Female
3 IU/ML 4.679535 1 wP Female
4 IU/ML 0.530000 3 wP Female
5 IU/ML 6.205949 3 wP Female
6 IU/ML 4.679535 3 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Unknown White 1983-01-01 2016-10-10 2020_dataset
5 Unknown White 1983-01-01 2016-10-10 2020_dataset
6 Unknown White 1983-01-01 2016-10-10 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1ggplot(igg) +
aes(MFI_normalised, antigen) +
geom_boxplot()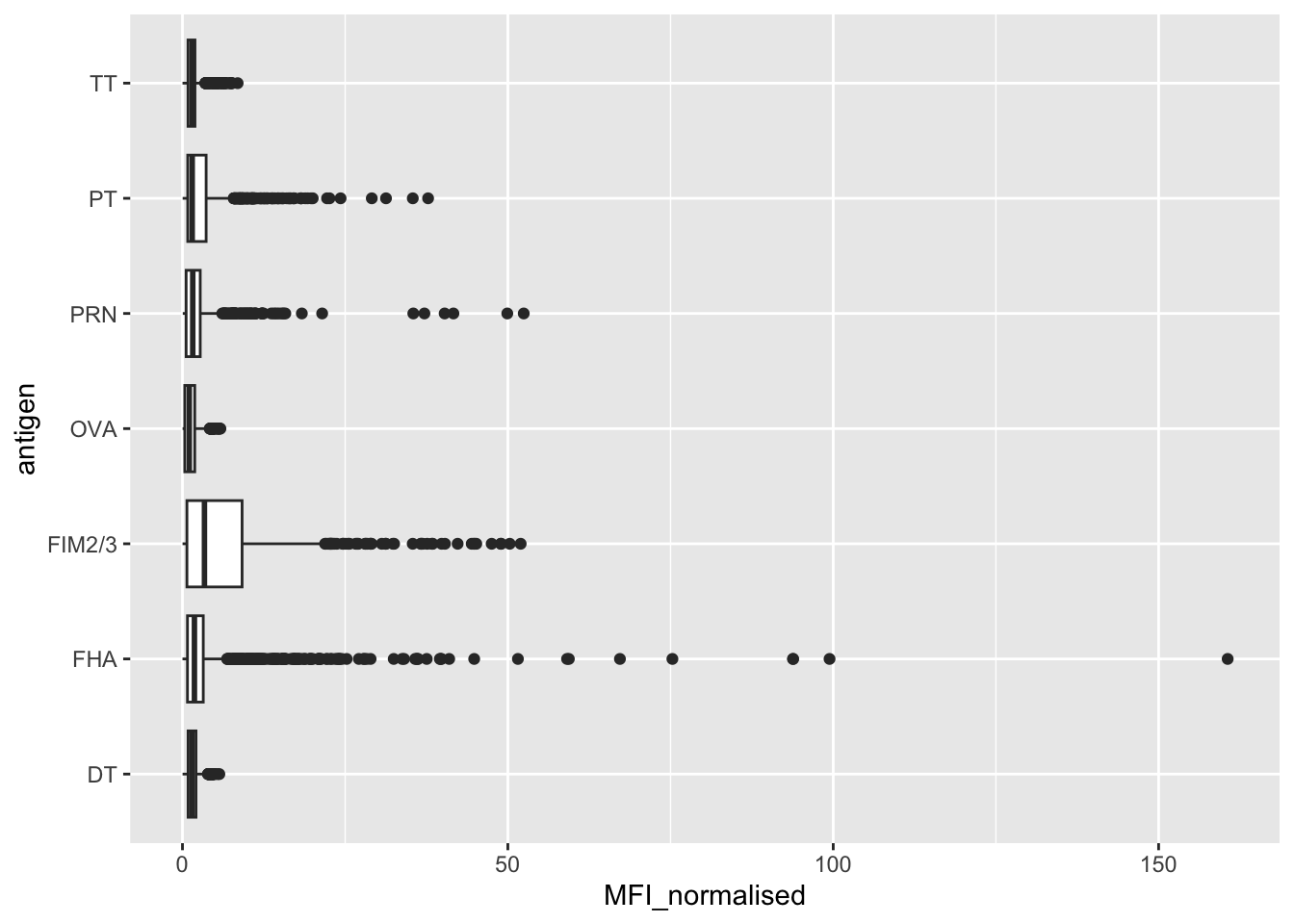
Q. Is there a difference for aP vs wP individuals with these values?
ggplot(igg) +
aes(MFI_normalised, antigen) +
geom_boxplot() +
facet_wrap(~infancy_vac)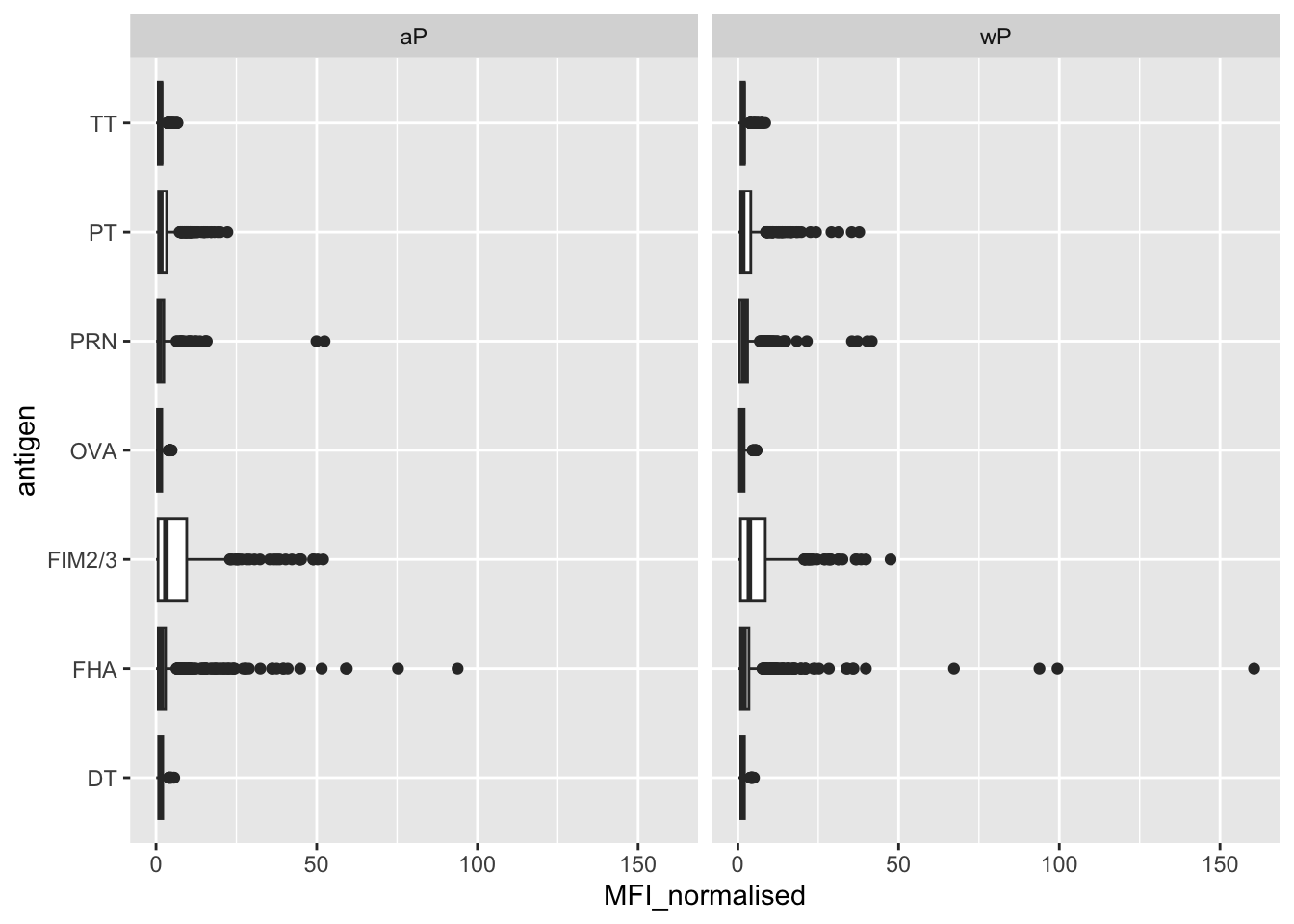
ggplot(igg) +
aes(MFI_normalised, antigen, col = infancy_vac) +
geom_boxplot()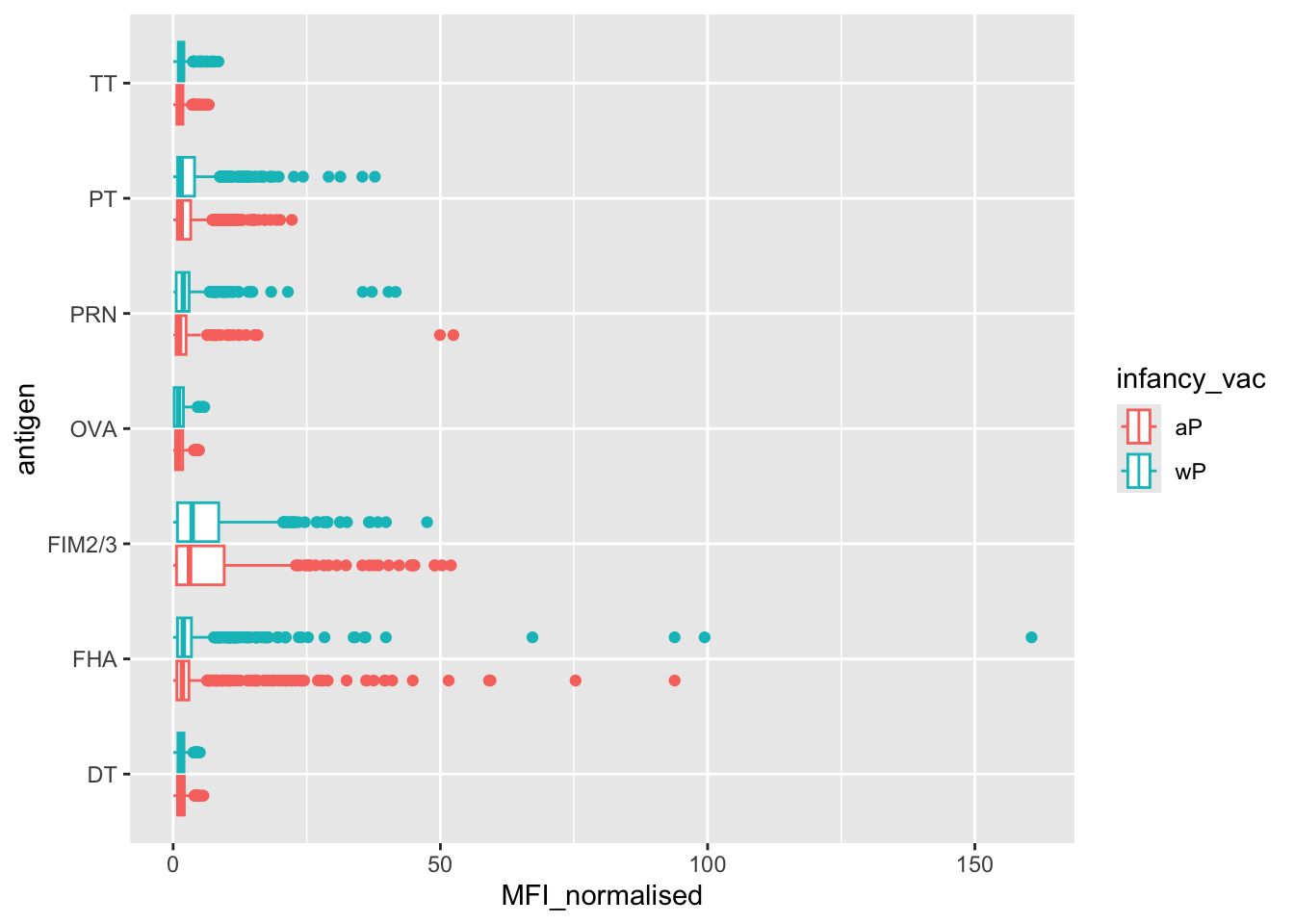
Q. Is there a temporal reponse - i.e. do these values increase or decease over time?
ggplot(igg) +
aes(MFI_normalised, antigen, col = infancy_vac) +
geom_boxplot() +
facet_wrap(~visit)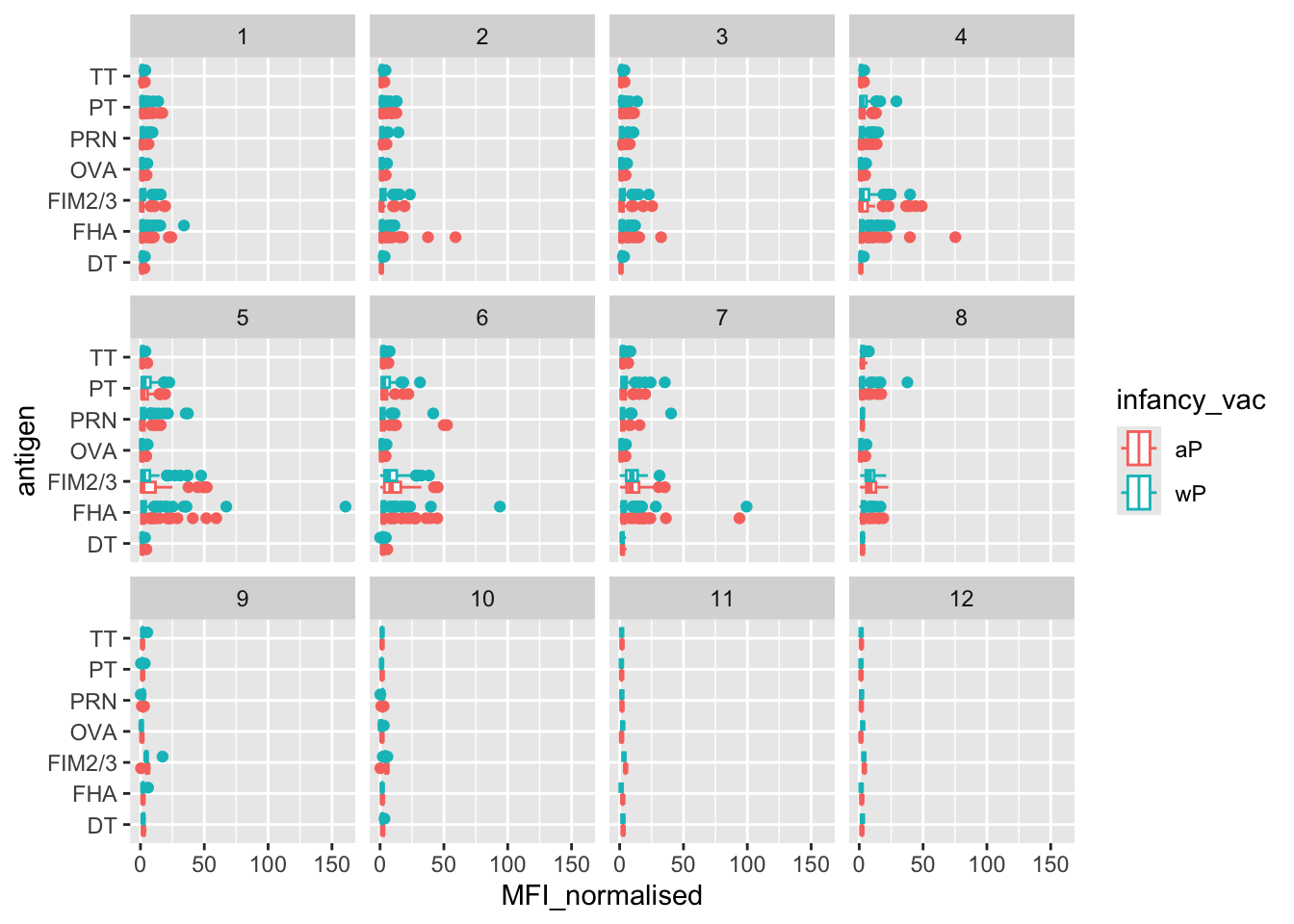
pt.igg.21 <- igg |> filter(antigen =="PT",
dataset == "2021_dataset")ggplot(pt.igg.21) +
aes(planned_day_relative_to_boost,
MFI_normalised,
col = infancy_vac,
group = subject_id) +
geom_point() +
geom_line() +
geom_vline(xintercept = 14, lty =2)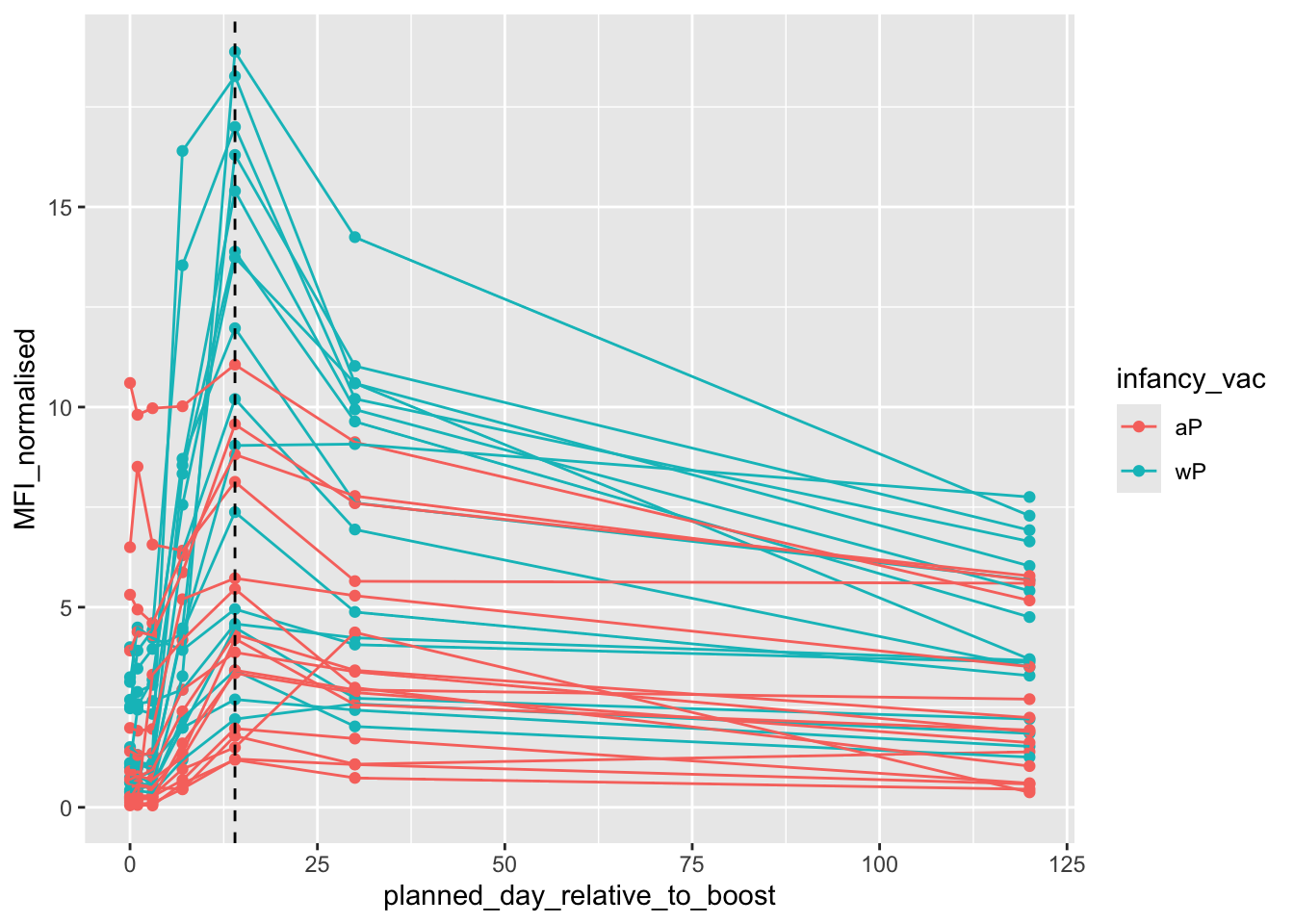
pt.igg.21 <- igg |>
filter(antigen == "PT", dataset == "2021_dataset")
pt.mean <- pt.igg.21 |>
group_by(infancy_vac, planned_day_relative_to_boost) |>
summarize(mean_MFI = mean(MFI_normalised, na.rm = TRUE), .groups = "drop")
pt.smooth <- pt.mean |>
group_by(infancy_vac) |>
group_modify(~{
xgrid <- seq(min(.x$planned_day_relative_to_boost),
max(.x$planned_day_relative_to_boost),
length.out = 200)
sf <- splinefun(.x$planned_day_relative_to_boost,
.x$mean_MFI,
method = "monoH.FC")
data.frame(
planned_day_relative_to_boost = xgrid,
mean_MFI = sf(xgrid)
)
})
ggplot() +
geom_point(data = pt.igg.21,
aes(x = planned_day_relative_to_boost,
y = MFI_normalised,
col = infancy_vac)) +
geom_line(data = pt.igg.21,
aes(x = planned_day_relative_to_boost,
y = MFI_normalised,
col = infancy_vac,
group = subject_id),
alpha = 0.25) +
geom_line(data = pt.smooth,
aes(x = planned_day_relative_to_boost,
y = mean_MFI,
col = infancy_vac),
linewidth = 1.8) +
geom_vline(xintercept = 0, linetype = "dashed") +
geom_vline(xintercept = 14, linetype = "dashed") +
labs(title = "CMI-PB 2021 dataset IgG PT",
subtitle = "Dashed lines at day 0 (pre boost) and day 14 (post boost)",
x = "Day relative to boost",
y = "Normalised MFI")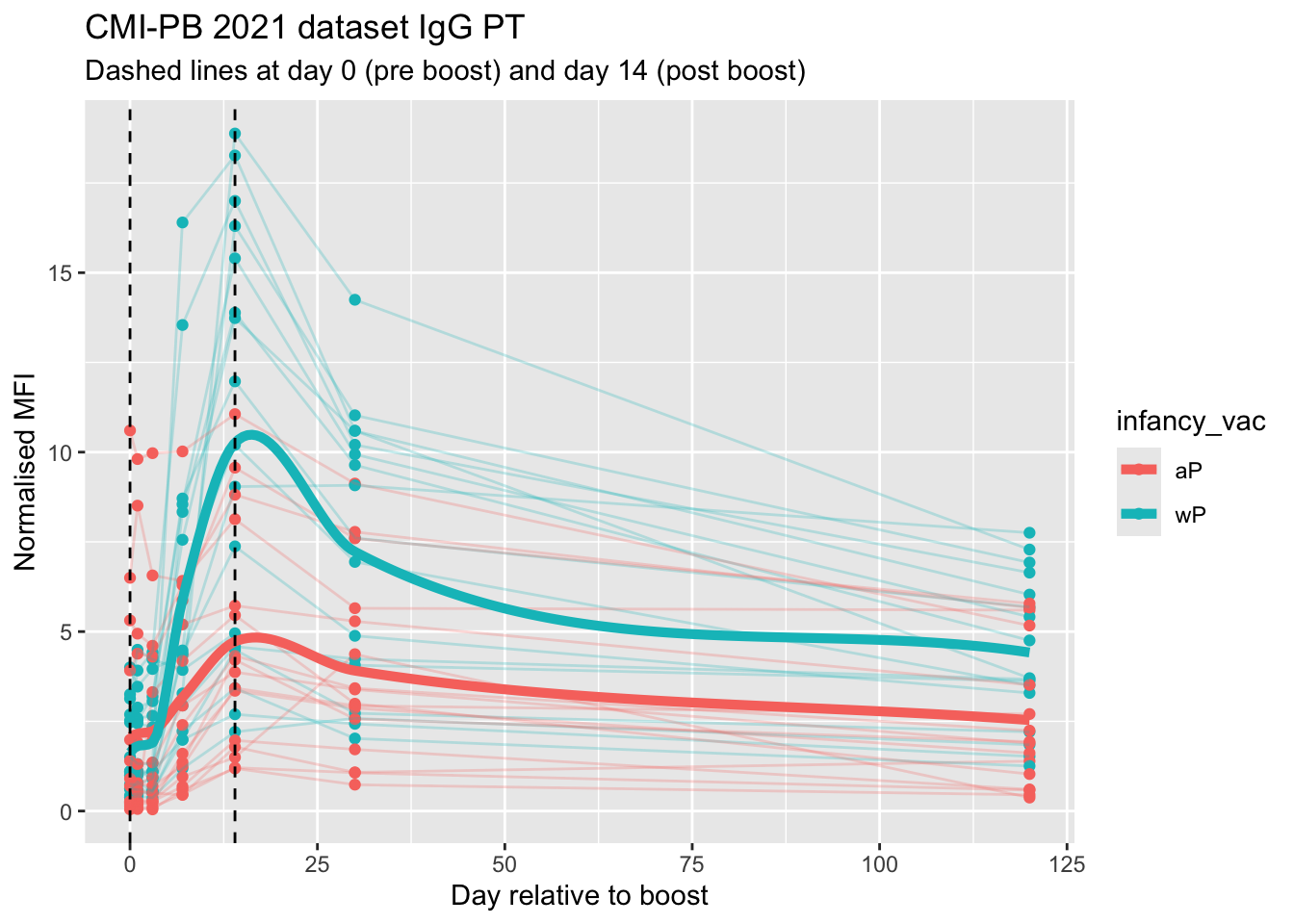
Q7. Using this approach determine (i) the average age of wP individuals, (ii) the average age of aP individuals; and (iii) are they significantly different?
library(lubridate)
library(dplyr)
subject$age <- today() - ymd(subject$year_of_birth)
ap <- subject %>% filter(infancy_vac == "aP")
wp <- subject %>% filter(infancy_vac == "wP")
mean_ap <- mean(time_length(ap$age, "years"))
mean_wp <- mean(time_length(wp$age, "years"))
mean_ap[1] 28.09932mean_wp[1] 36.85002t.test(time_length(wp$age, "years"),
time_length(ap$age, "years"))
Welch Two Sample t-test
data: time_length(wp$age, "years") and time_length(ap$age, "years")
t = 12.918, df = 104.03, p-value < 2.2e-16
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
7.407351 10.094058
sample estimates:
mean of x mean of y
36.85002 28.09932 The wP group on average is older than the aP group. From this Welch Two Sample t-test the age difference betwen the two groups is definitey significant.
Q8. Determine the age of all individuals at time of boost?
subject$age_at_boost <- time_length(
ymd(subject$date_of_boost) - ymd(subject$year_of_birth),
"years"
)
head(subject$age_at_boost)[1] 30.69678 51.07461 33.77413 28.65982 25.65914 28.77481summary(subject$age_at_boost) Min. 1st Qu. Median Mean 3rd Qu. Max.
18.83 21.03 25.75 26.09 29.56 51.07 In this table, we can view the age of all individuals at the time of the booster vaccine.
Q9. With the help of a faceted boxplot or histogram (see below), do you think these two groups are significantly different?
ggplot(subject) +
aes(time_length(age, "year"),
fill = as.factor(infancy_vac)) +
geom_histogram(show.legend = FALSE) +
facet_wrap(vars(infancy_vac), nrow = 2) +
xlab("Age in years")`stat_bin()` using `bins = 30`. Pick better value `binwidth`.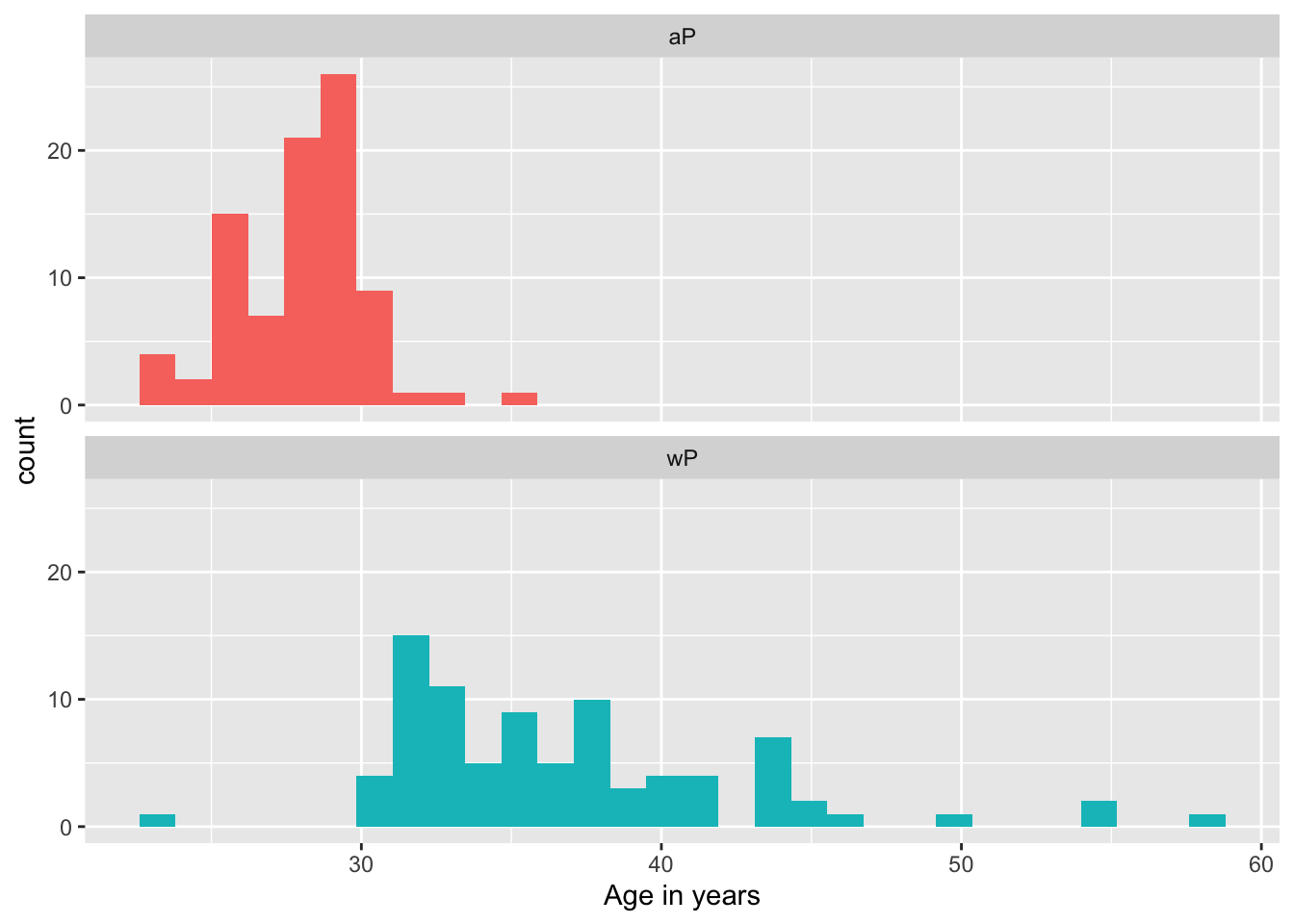
Yes, these two age groups are significantly different. We can see that the wP group is shifted older the aP group. This is as expected due to the vaccine switch.
Q10. Now using the same procedure join meta with titer data so we can further analyze this data in terms of time of visit aP/wP, male/female etc.
head(meta) subject_id infancy_vac biological_sex ethnicity race
1 1 wP Female Not Hispanic or Latino White
2 1 wP Female Not Hispanic or Latino White
3 1 wP Female Not Hispanic or Latino White
4 1 wP Female Not Hispanic or Latino White
5 1 wP Female Not Hispanic or Latino White
6 1 wP Female Not Hispanic or Latino White
year_of_birth date_of_boost dataset specimen_id
1 1986-01-01 2016-09-12 2020_dataset 1
2 1986-01-01 2016-09-12 2020_dataset 2
3 1986-01-01 2016-09-12 2020_dataset 3
4 1986-01-01 2016-09-12 2020_dataset 4
5 1986-01-01 2016-09-12 2020_dataset 5
6 1986-01-01 2016-09-12 2020_dataset 6
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 1 1 Blood
3 3 3 Blood
4 7 7 Blood
5 11 14 Blood
6 32 30 Blood
visit
1 1
2 2
3 3
4 4
5 5
6 6head(abdata) specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgE FALSE Total 1110.21154 2.493425
2 1 IgE FALSE Total 2708.91616 2.493425
3 1 IgG TRUE PT 68.56614 3.736992
4 1 IgG TRUE PRN 332.12718 2.602350
5 1 IgG TRUE FHA 1887.12263 34.050956
6 1 IgE TRUE ACT 0.10000 1.000000
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 UG/ML 2.096133 1 wP Female
2 IU/ML 29.170000 1 wP Female
3 IU/ML 0.530000 1 wP Female
4 IU/ML 6.205949 1 wP Female
5 IU/ML 4.679535 1 wP Female
6 IU/ML 2.816431 1 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
5 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
6 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1Q11. How many specimens (i.e. entries in abdata) do we have for each isotype?
table(abdata$isotype)
IgE IgG IgG1 IgG2 IgG3 IgG4
6698 7265 11993 12000 12000 12000 Q12. What are the different $dataset values in abdata and what do you notice about the number of rows for the most “recent” dataset?
table(abdata$dataset)
2020_dataset 2021_dataset 2022_dataset 2023_dataset
31520 8085 7301 15050 This dataset column contains many study datasets, which include the 2020_dataset and 2021_dataset. I noticed that the most recent dataset has less rows than the older ones, which could indicate fewer measurements or subjects.
igg <- abdata %>% filter(isotype == "IgG")
head(igg) specimen_id isotype is_antigen_specific antigen MFI MFI_normalised
1 1 IgG TRUE PT 68.56614 3.736992
2 1 IgG TRUE PRN 332.12718 2.602350
3 1 IgG TRUE FHA 1887.12263 34.050956
4 19 IgG TRUE PT 20.11607 1.096366
5 19 IgG TRUE PRN 976.67419 7.652635
6 19 IgG TRUE FHA 60.76626 1.096457
unit lower_limit_of_detection subject_id infancy_vac biological_sex
1 IU/ML 0.530000 1 wP Female
2 IU/ML 6.205949 1 wP Female
3 IU/ML 4.679535 1 wP Female
4 IU/ML 0.530000 3 wP Female
5 IU/ML 6.205949 3 wP Female
6 IU/ML 4.679535 3 wP Female
ethnicity race year_of_birth date_of_boost dataset
1 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
2 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
3 Not Hispanic or Latino White 1986-01-01 2016-09-12 2020_dataset
4 Unknown White 1983-01-01 2016-10-10 2020_dataset
5 Unknown White 1983-01-01 2016-10-10 2020_dataset
6 Unknown White 1983-01-01 2016-10-10 2020_dataset
actual_day_relative_to_boost planned_day_relative_to_boost specimen_type
1 -3 0 Blood
2 -3 0 Blood
3 -3 0 Blood
4 -3 0 Blood
5 -3 0 Blood
6 -3 0 Blood
visit
1 1
2 1
3 1
4 1
5 1
6 1Q13. Complete the following code to make a summary boxplot of Ab titer levels (MFI) for all antigens:
ggplot(igg) +
aes(MFI_normalised, antigen) +
geom_boxplot() +
xlim(0, 75) +
facet_wrap(vars(visit), nrow = 2)Warning: Removed 5 rows containing non-finite outside the scale range
(`stat_boxplot()`).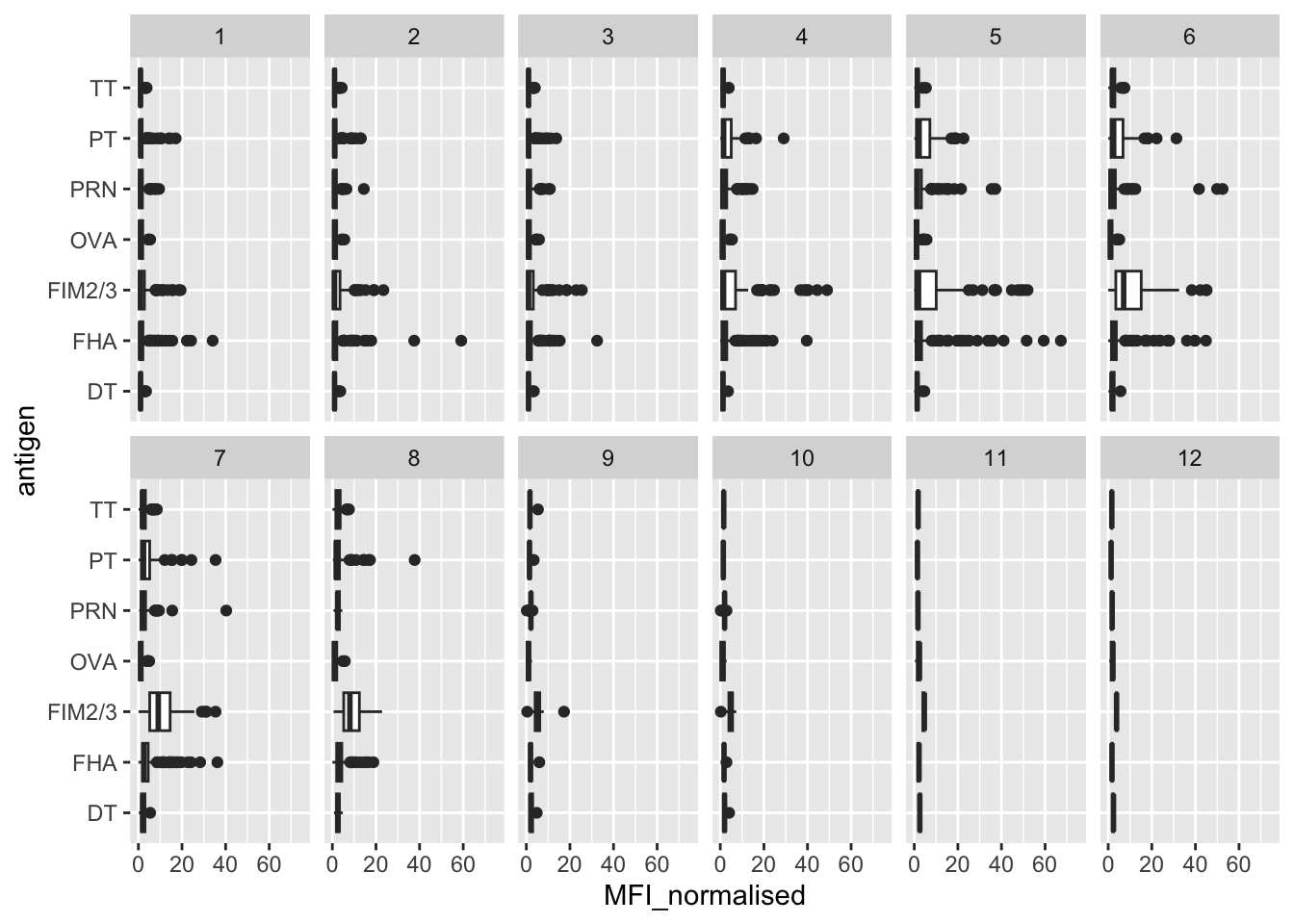
Q14. What antigens show differences in the level of IgG antibody titers recognizing them over time? Why these and not others?
The antigens that show the differences are PT, PRN, FHA, and FIM2/3. These antigens appear to rise after a booster and then fall again. A control antigen like the OVA is one that appears to stay low over time since it is not part of the vaccine response.
Q15. Filter to pull out only two specific antigens for analysis and create a boxplot for each. You can chose any you like. Below I picked a “control” antigen (“OVA”, that is not in our vaccines) and a clear antigen of interest (“PT”, Pertussis Toxin, one of the key virulence factors produced by the bacterium B. pertussis).
filter(igg, antigen == "OVA") %>%
ggplot() +
aes(MFI_normalised, col = infancy_vac) +
geom_boxplot(show.legend = FALSE) +
facet_wrap(vars(visit)) +
theme_bw()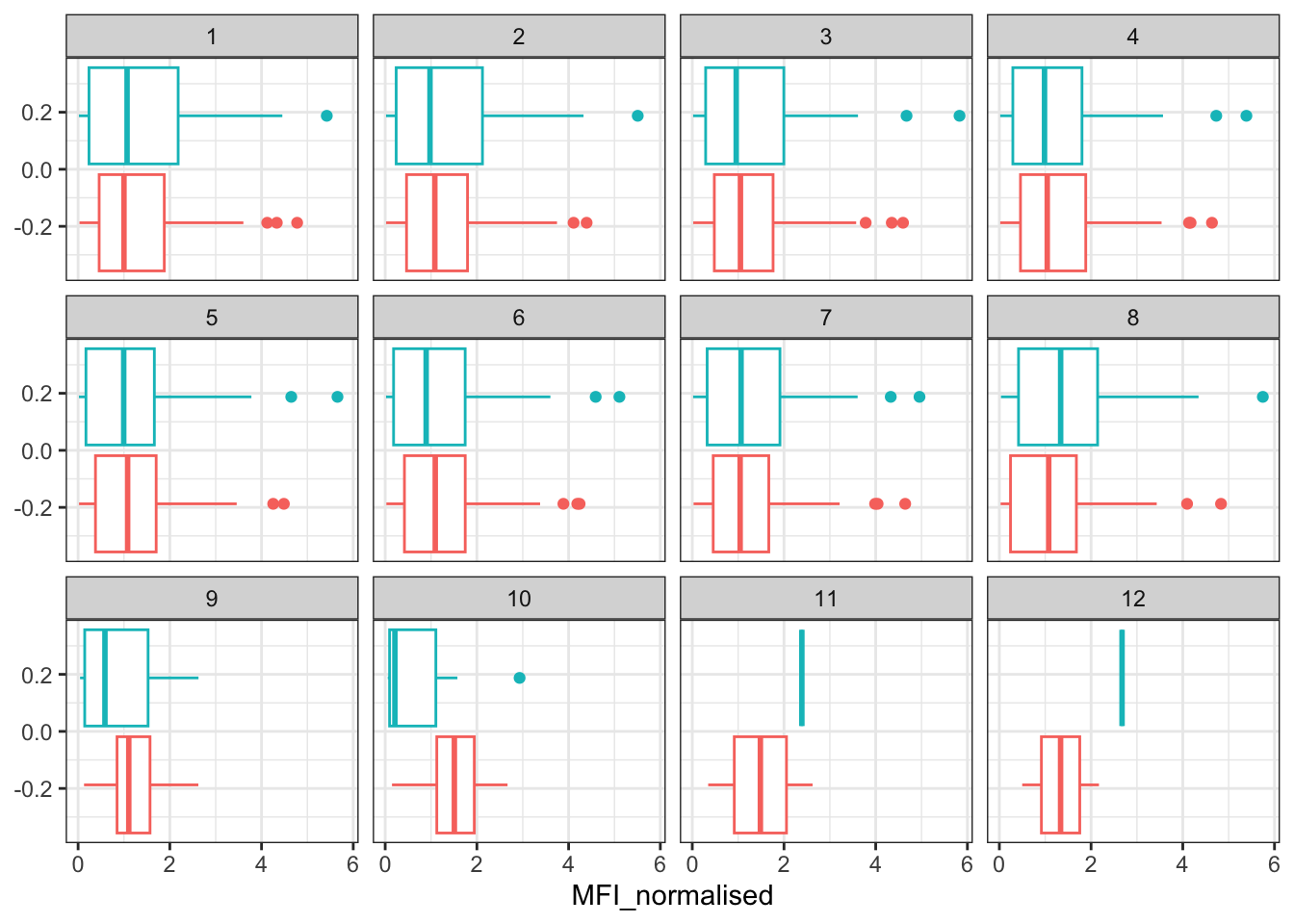
filter(igg, antigen == "PT") %>%
ggplot() +
aes(MFI_normalised, col = infancy_vac) +
geom_boxplot(show.legend = FALSE) +
facet_wrap(vars(visit)) +
theme_bw()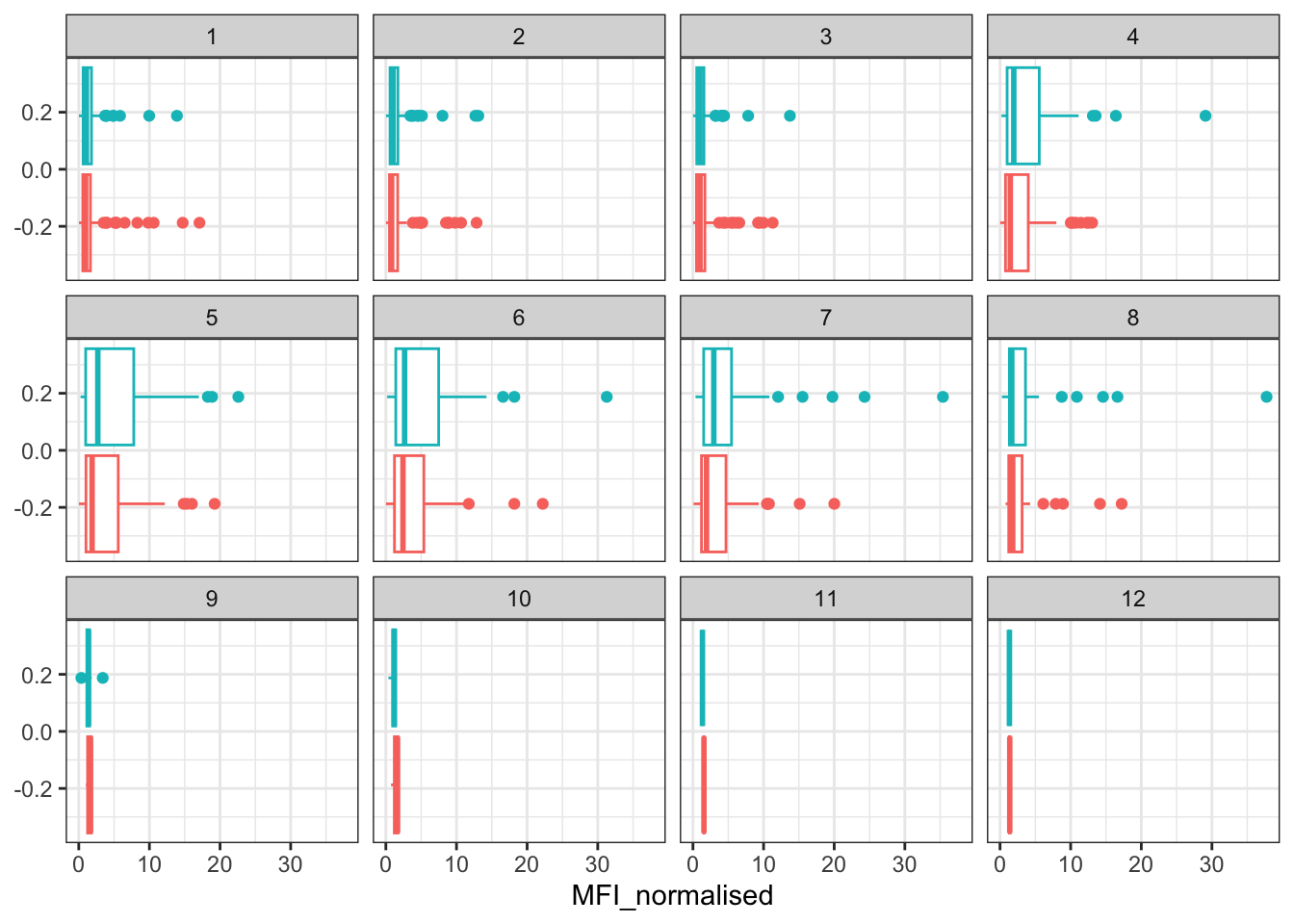
Q16. What do you notice about these two antigens time courses and the PT data in particular?
I noticed that the OVA antigen acts like a negative control and seems to stay relatively low over time with low evidence of a strong booster response. The PT antigen shows more evidence of a rise after the booster and peaks around the middle visits, then declines afterwards. PT seems to be the one with the expected antigen-specific immunse reponse.
Q17. Do you see any clear difference in aP vs. wP responses?
There seems to be some differences between aP and wP responses. The wP response is higher for pertussus related responses but there is also some overlap between the groups.
abdata.21 <- abdata %>% filter(dataset == "2021_dataset")
abdata.21 %>%
filter(isotype == "IgG", antigen == "PT") %>%
ggplot() +
aes(x=planned_day_relative_to_boost,
y=MFI_normalised,
col=infancy_vac,
group=subject_id) +
geom_point() +
geom_line() +
geom_vline(xintercept=0, linetype="dashed") +
geom_vline(xintercept=14, linetype="dashed") +
labs(title="2021 dataset IgG PT",
subtitle = "Dashed lines indicate day 0 (pre-boost) and 14 (apparent peak levels)")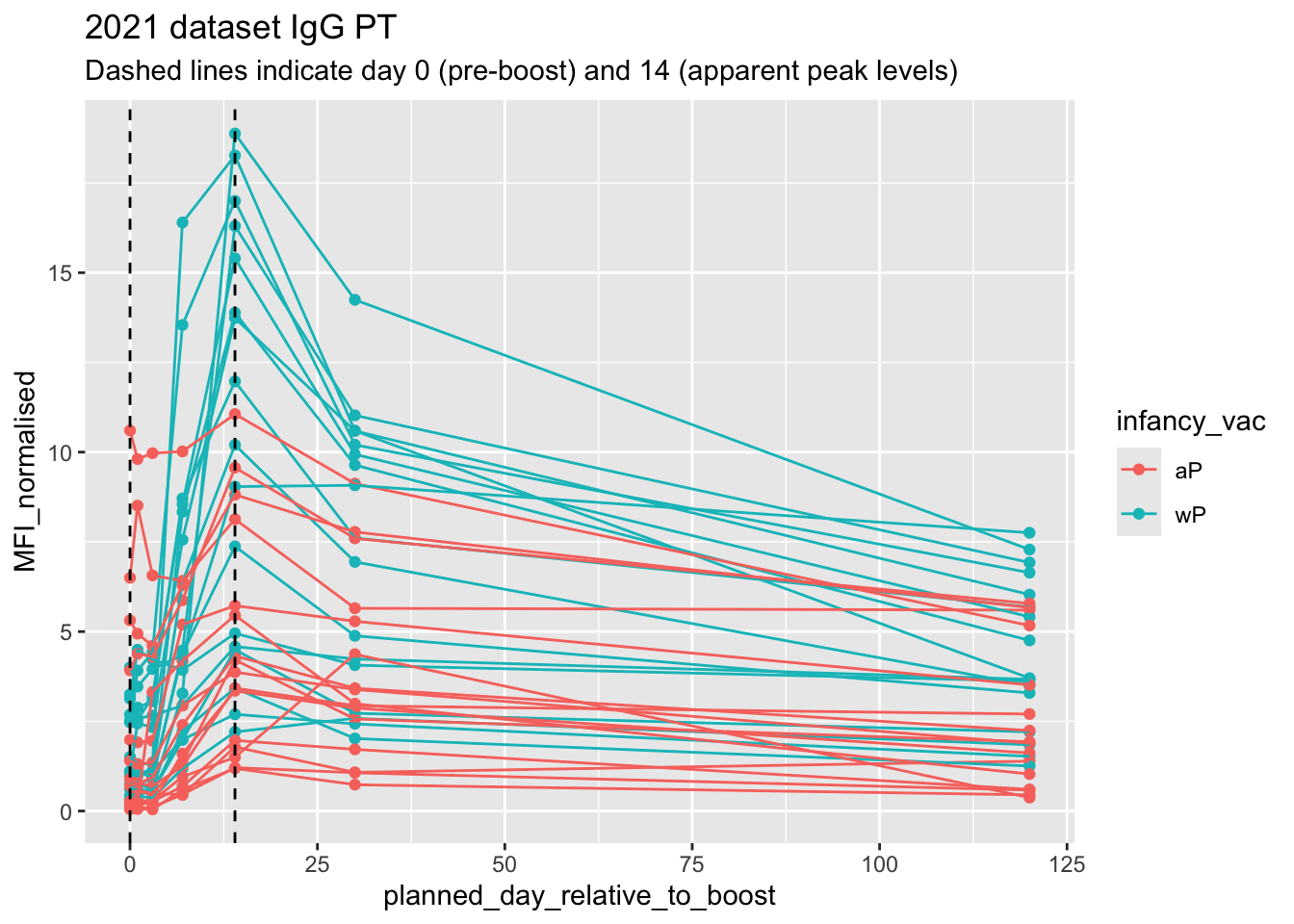
Q18. Does this trend look similar for the 2020 dataset?
Yes, this trend looks similar for the 2020 dataset. We can see that antibody levels rise after the boost and peak around the earlier/middle boost period and then begin to decline.
rna_url <- "https://www.cmi-pb.org/api/v2/rnaseq?versioned_ensembl_gene_id=eq.ENSG00000211896.7"
if (!file.exists("IGHG1_rna.json")) {
download.file(rna_url, destfile = "IGHG1_rna.json", mode = "wb")
}
rna <- read_json("IGHG1_rna.json", simplifyVector = TRUE)
ssrna <- inner_join(rna, meta, by = "specimen_id")Q19. Make a plot of the time course of gene expression for IGHG1 gene (i.e. a plot of visit vs. tpm).
ggplot(ssrna) +
aes(visit, tpm, group = subject_id) +
geom_point() +
geom_line(alpha = 0.2)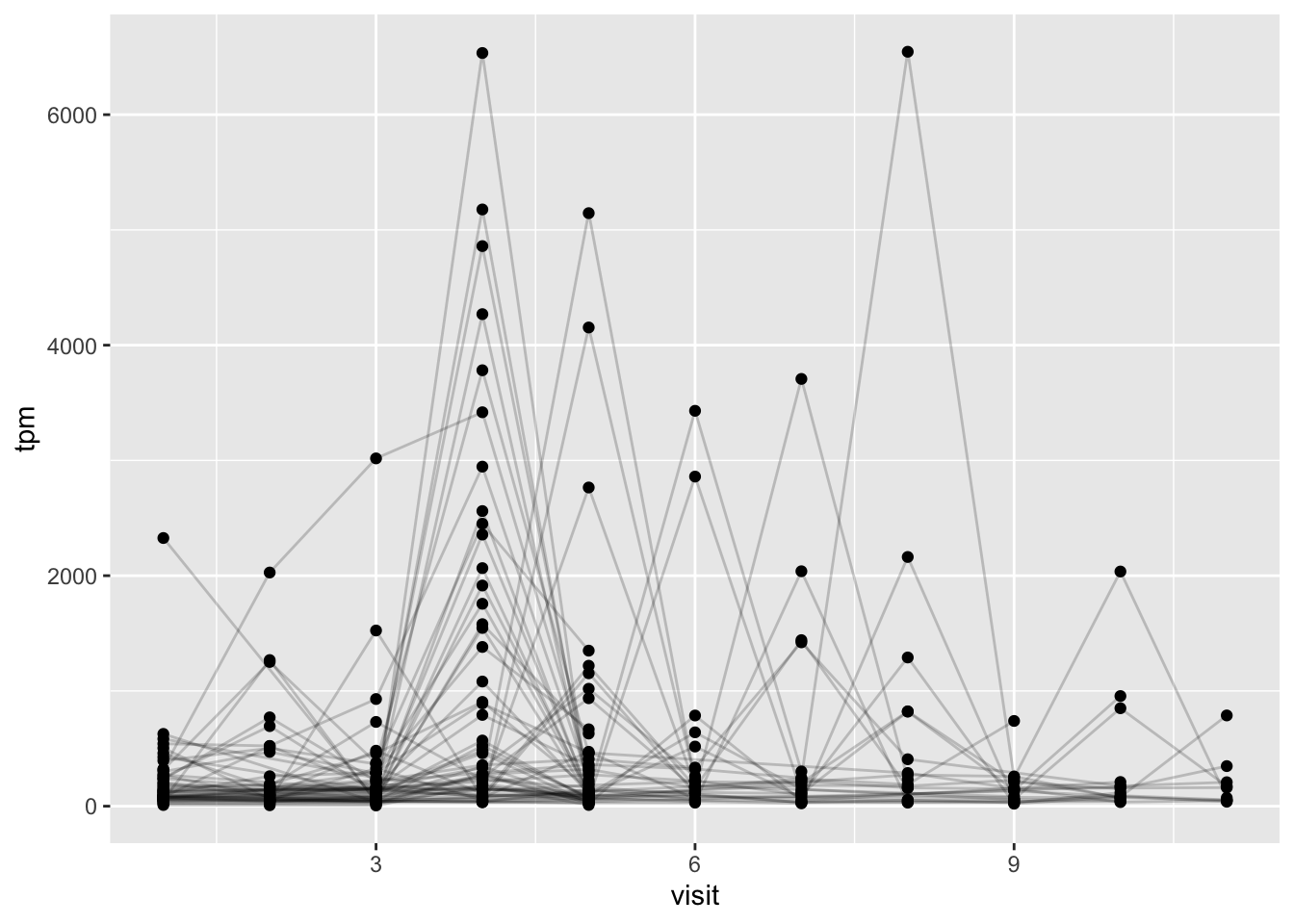
Q20.: What do you notice about the expression of this gene (i.e. when is it at it’s maximum level)?
This IGHG1 gene expression seems to be highest around the time the booster is administered and around the early post-boost visits instead of later visits.
- Does this pattern in time match the trend of antibody titer data? If not, why not?
This pattern doesn’t fully match the trend of antibody titer data because the gene expression changes occur earlier than antibody titer changes. Titer peaks later and stays elevated for a longer period of time.
We can dig deeper and color and/or facet by infancy_vac status:
ggplot(ssrna) +
aes(tpm, col=infancy_vac) +
geom_boxplot() +
facet_wrap(vars(visit))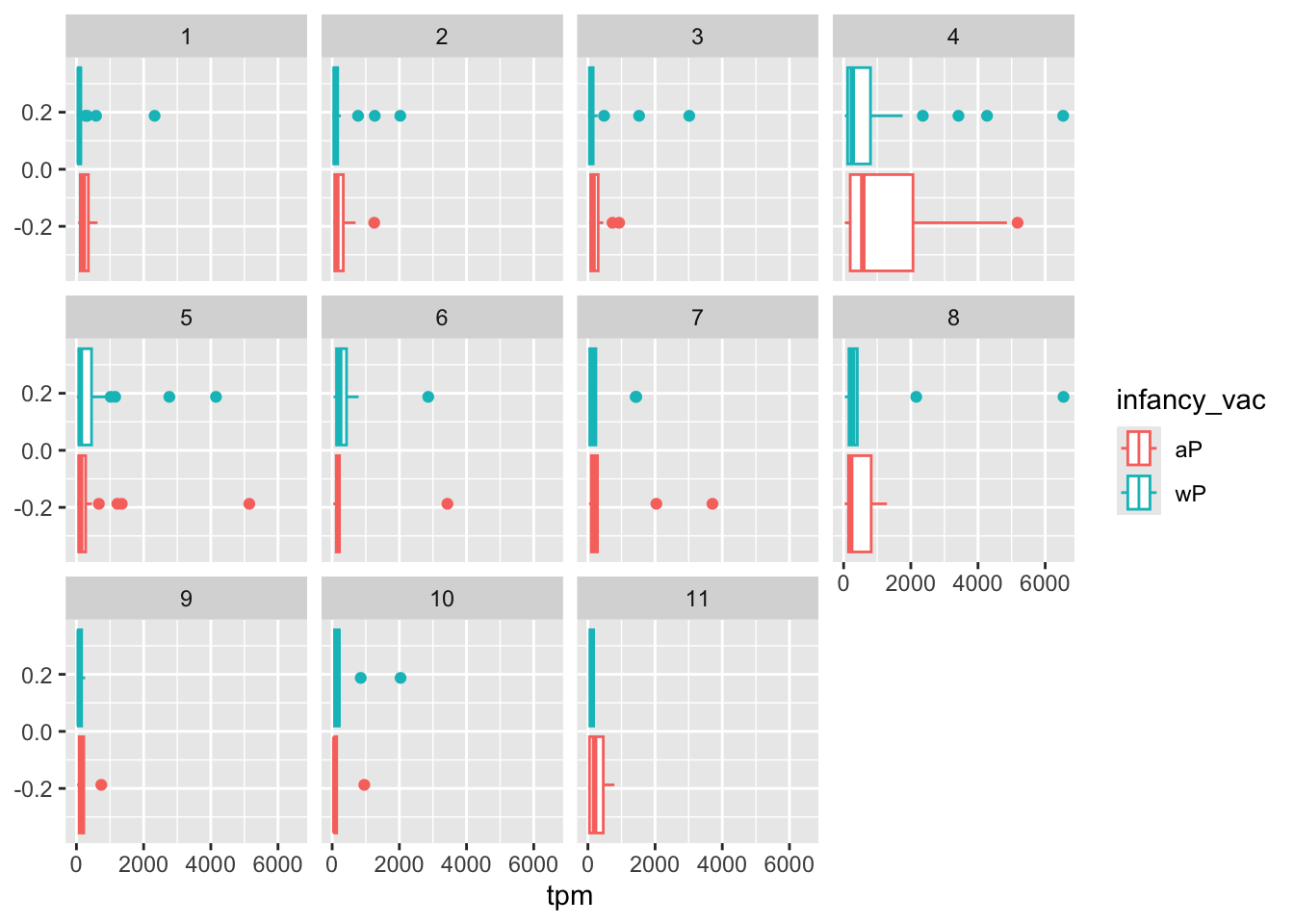
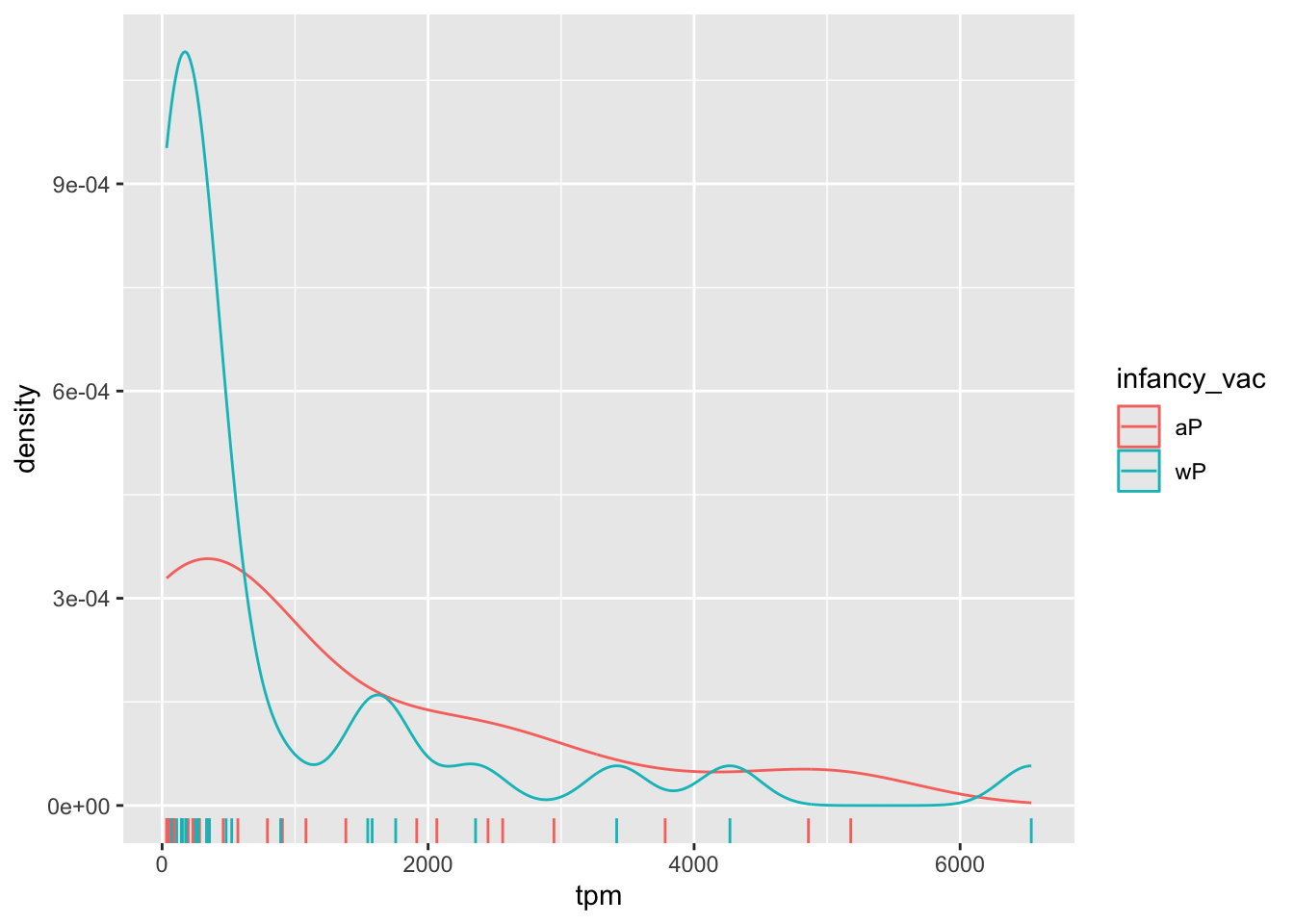
There is however no obvious wP vs. aP differences here even if we focus in on a particular visit:
ssrna %>%
filter(visit==4) %>%
ggplot() +
aes(tpm, col=infancy_vac) + geom_density() +
geom_rug()