(250000 / 200000000) *100[1] 0.125In this hands-on session we will utilize AlphaFold to predict protein structure from sequence (Jumper et al. 2021).
Without the aid of such approaches, it can take years of expensive laboratory work to determine the structure of just one protein. With AlphaFold we can now accurately compute a typical protein structure in as little as ten minutes.
The PDB database (the main repository of experimental structures) only has ~250 thousand structures (we saw this in the last lab). The main protein sequence database has over 200 million sequences! Only 0.125% of known sequences have a known structure - this is called the “structure knowledge gap”.
(250000 / 200000000) *100[1] 0.125We can download and run locally (on our own computers) but we need a GPU. Or we can use “cloud” computing to run this on someone elses computers.
We will use ColabFold https://github.com/sokrypton/ColabFold
We previously found there was no AFDB entry for our HIV sequence:
>HIV-Pr-Dimer
PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIGGFIKVRQYD
QILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF:PQITLWQRPLVTIKIGGQLK
EALLDTGADDTVLEEMSLPGRWKPKMIGGIGGFIKVRQYDQILIEICGHKAIGTVLVGPT
PVNIIGRNLLTQIGCTLNFHere we will use AlphaFold2_mmseqs2
After I ran this in the online GPU resource, ColabFold get back a downloadable zip file that contained five ranked structure models in PDB format, MSA alignment files, and JSON files.
I unzipped the downloadable files and looked in Mol to see a visualization and also in Bio3D.
library(bio3d)
library(jsonlite)
library(pheatmap)
results_dir <- "hivpr_23119/"
list.files(results_dir)[1:20] [1] "cite.bibtex"
[2] "config.json"
[3] "hivpr_23119_coverage.png"
[4] "hivpr_23119_env"
[5] "hivpr_23119_pae.png"
[6] "hivpr_23119_plddt.png"
[7] "hivpr_23119_predicted_aligned_error_v1.json"
[8] "hivpr_23119_scores_rank_001_alphafold2_multimer_v3_model_4_seed_000.json"
[9] "hivpr_23119_scores_rank_002_alphafold2_multimer_v3_model_1_seed_000.json"
[10] "hivpr_23119_scores_rank_003_alphafold2_multimer_v3_model_5_seed_000.json"
[11] "hivpr_23119_scores_rank_004_alphafold2_multimer_v3_model_2_seed_000.json"
[12] "hivpr_23119_scores_rank_005_alphafold2_multimer_v3_model_3_seed_000.json"
[13] "hivpr_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb"
[14] "hivpr_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb"
[15] "hivpr_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb"
[16] "hivpr_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb"
[17] "hivpr_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb"
[18] "hivpr_23119.a3m"
[19] "hivpr_23119.csv"
[20] "hivpr_23119.done.txt" library(bio3d)
pdb_files <- list.files(path = results_dir,
pattern = "\\.pdb$",
full.names = TRUE)
basename(pdb_files)[1] "hivpr_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb"
[2] "hivpr_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb"
[3] "hivpr_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb"
[4] "hivpr_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb"
[5] "hivpr_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb"length(pdb_files)[1] 5pdbs <- pdbaln(pdb_files, fit = TRUE, exefile = "msa")Reading PDB files:
hivpr_23119//hivpr_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb
hivpr_23119//hivpr_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb
hivpr_23119//hivpr_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb
hivpr_23119//hivpr_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb
hivpr_23119//hivpr_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb
.....
Extracting sequences
pdb/seq: 1 name: hivpr_23119//hivpr_23119_unrelaxed_rank_001_alphafold2_multimer_v3_model_4_seed_000.pdb
pdb/seq: 2 name: hivpr_23119//hivpr_23119_unrelaxed_rank_002_alphafold2_multimer_v3_model_1_seed_000.pdb
pdb/seq: 3 name: hivpr_23119//hivpr_23119_unrelaxed_rank_003_alphafold2_multimer_v3_model_5_seed_000.pdb
pdb/seq: 4 name: hivpr_23119//hivpr_23119_unrelaxed_rank_004_alphafold2_multimer_v3_model_2_seed_000.pdb
pdb/seq: 5 name: hivpr_23119//hivpr_23119_unrelaxed_rank_005_alphafold2_multimer_v3_model_3_seed_000.pdb pdbs 1 . . . . 50
[Truncated_Name:1]hivpr_2311 PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGI
[Truncated_Name:2]hivpr_2311 PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGI
[Truncated_Name:3]hivpr_2311 PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGI
[Truncated_Name:4]hivpr_2311 PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGI
[Truncated_Name:5]hivpr_2311 PQITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGI
**************************************************
1 . . . . 50
51 . . . . 100
[Truncated_Name:1]hivpr_2311 GGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNFP
[Truncated_Name:2]hivpr_2311 GGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNFP
[Truncated_Name:3]hivpr_2311 GGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNFP
[Truncated_Name:4]hivpr_2311 GGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNFP
[Truncated_Name:5]hivpr_2311 GGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNFP
**************************************************
51 . . . . 100
101 . . . . 150
[Truncated_Name:1]hivpr_2311 QITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIG
[Truncated_Name:2]hivpr_2311 QITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIG
[Truncated_Name:3]hivpr_2311 QITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIG
[Truncated_Name:4]hivpr_2311 QITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIG
[Truncated_Name:5]hivpr_2311 QITLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIG
**************************************************
101 . . . . 150
151 . . . . 198
[Truncated_Name:1]hivpr_2311 GFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
[Truncated_Name:2]hivpr_2311 GFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
[Truncated_Name:3]hivpr_2311 GFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
[Truncated_Name:4]hivpr_2311 GFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
[Truncated_Name:5]hivpr_2311 GFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
************************************************
151 . . . . 198
Call:
pdbaln(files = pdb_files, fit = TRUE, exefile = "msa")
Class:
pdbs, fasta
Alignment dimensions:
5 sequence rows; 198 position columns (198 non-gap, 0 gap)
+ attr: xyz, resno, b, chain, id, ali, resid, sse, callRMSD (heatmap)
library(pheatmap)
rd <- rmsd(pdbs, fit = TRUE)Warning in rmsd(pdbs, fit = TRUE): No indices provided, using the 198 non NA positionsrange(rd)[1] 0.000 14.754colnames(rd) <- paste0("m", 1:nrow(rd))
rownames(rd) <- paste0("m", 1:nrow(rd))
pheatmap(rd)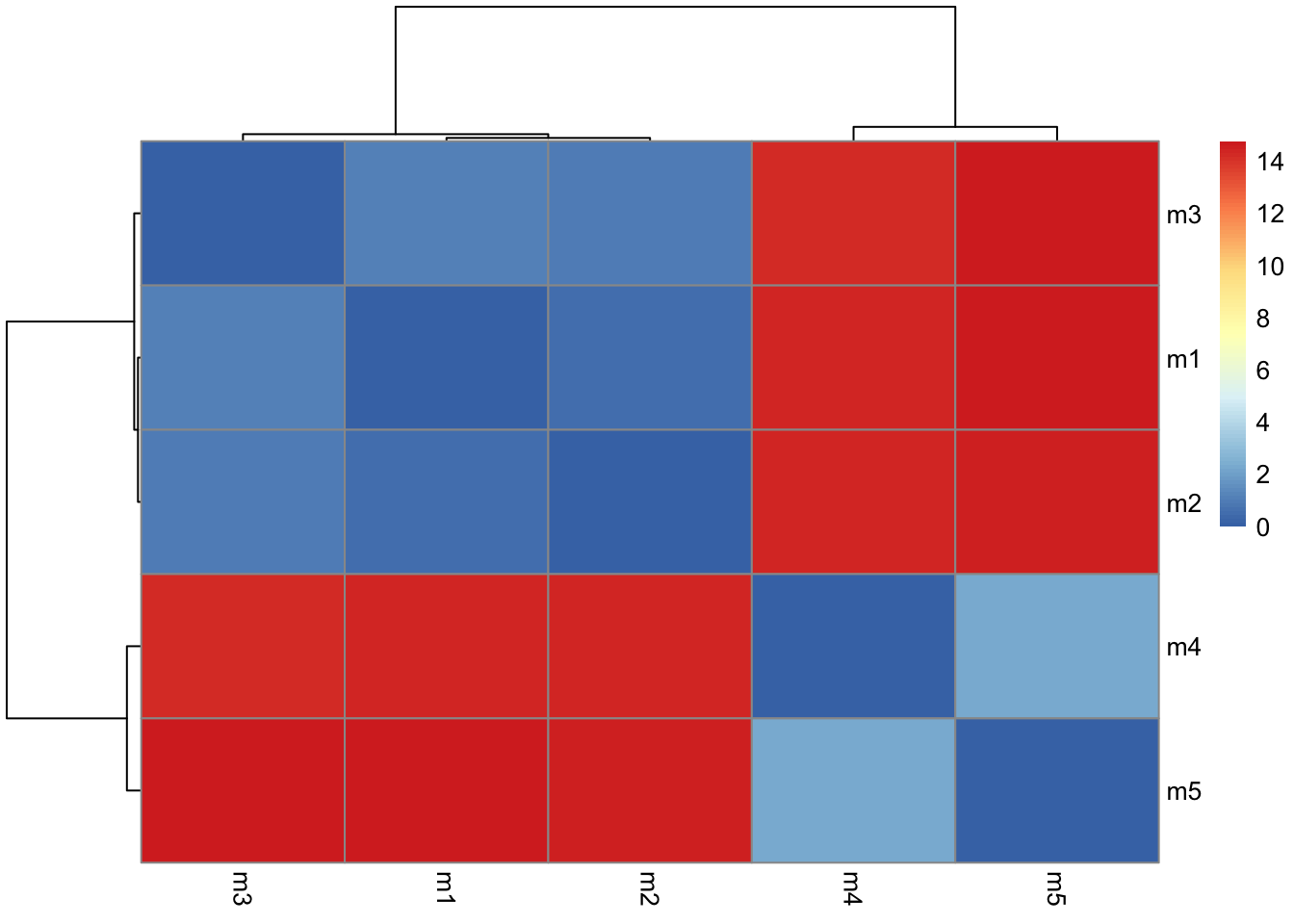
From our RMSD range, we can see that certain models were very similar while others were very different from each other.
pdb_ref <- read.pdb("1hsg") Note: Accessing on-line PDB fileplotb3(pdbs$b[1,], typ="l", lwd=2, sse=pdb_ref,
ylab="pLDDT (B-factor)", xlab="Residue Position")
lines(pdbs$b[2,], col="red")
lines(pdbs$b[3,], col="blue")
lines(pdbs$b[4,], col="darkgreen")
lines(pdbs$b[5,], col="orange")
abline(v = 99, col = "gray")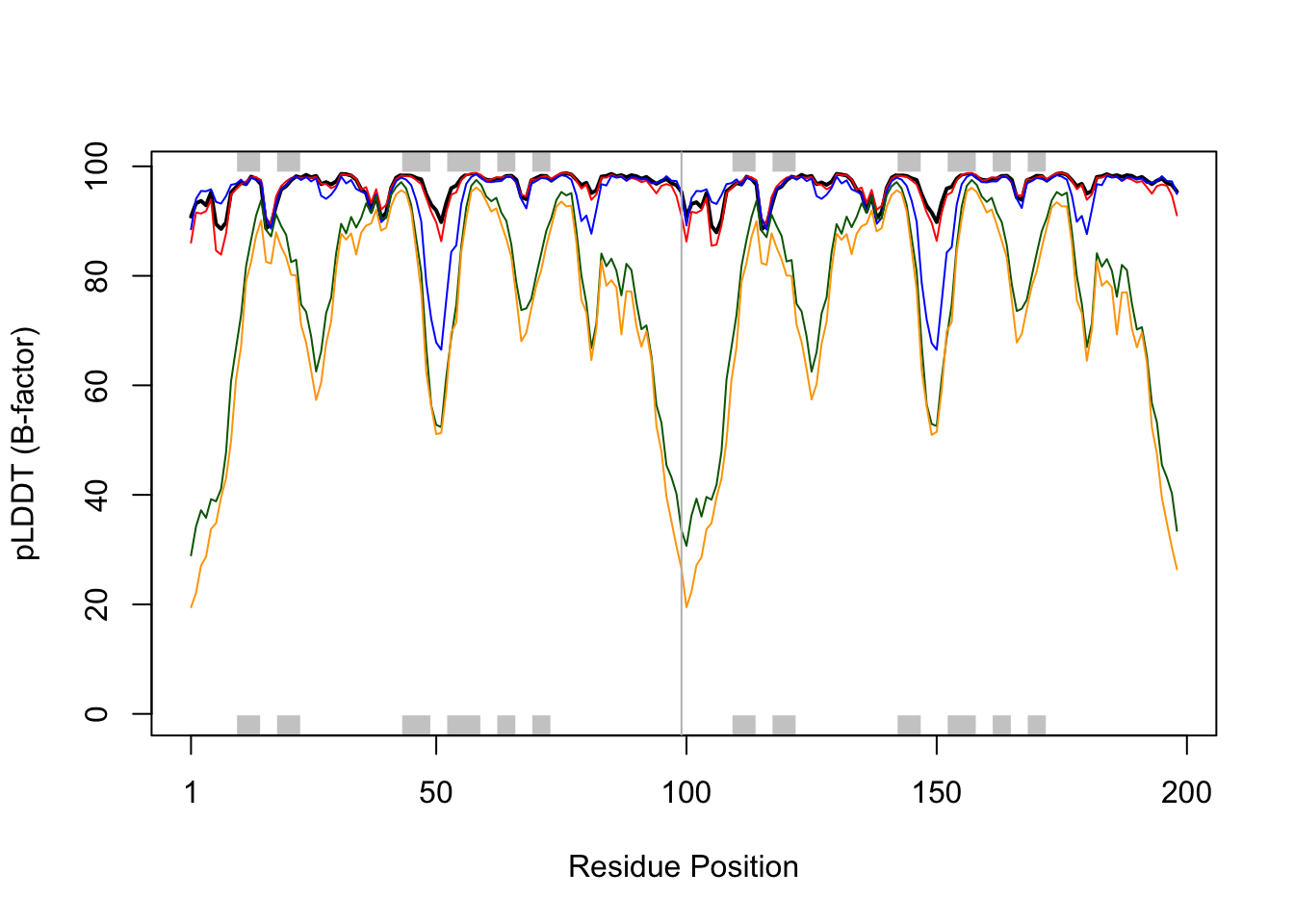
library(jsonlite)
pae_files <- list.files(path = results_dir,
pattern = "scores_rank_.*\\.json$",
full.names = TRUE)
basename(pae_files)[1] "hivpr_23119_scores_rank_001_alphafold2_multimer_v3_model_4_seed_000.json"
[2] "hivpr_23119_scores_rank_002_alphafold2_multimer_v3_model_1_seed_000.json"
[3] "hivpr_23119_scores_rank_003_alphafold2_multimer_v3_model_5_seed_000.json"
[4] "hivpr_23119_scores_rank_004_alphafold2_multimer_v3_model_2_seed_000.json"
[5] "hivpr_23119_scores_rank_005_alphafold2_multimer_v3_model_3_seed_000.json"pae1 <- read_json(pae_files[1], simplifyVector = TRUE)
pae5 <- read_json(pae_files[5], simplifyVector = TRUE)
pae1$max_pae[1] 12.84375pae5$max_pae[1] 29.59375plot.dmat(pae1$pae, xlab="Residue i", ylab="Residue j",
grid.col="black", zlim=c(0,30))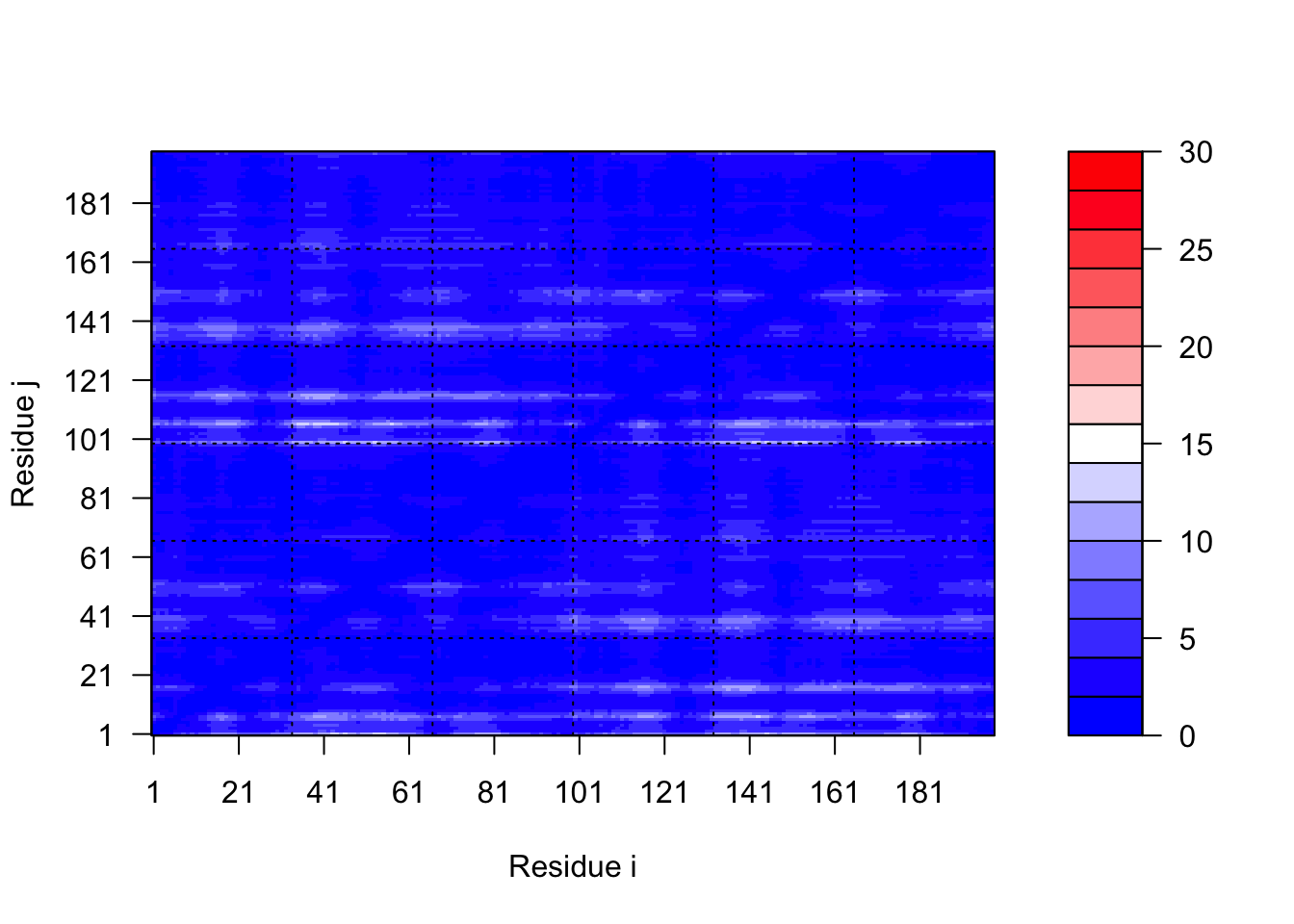
plot.dmat(pae5$pae, xlab="Residue i", ylab="Residue j",
grid.col="black", zlim=c(0,30))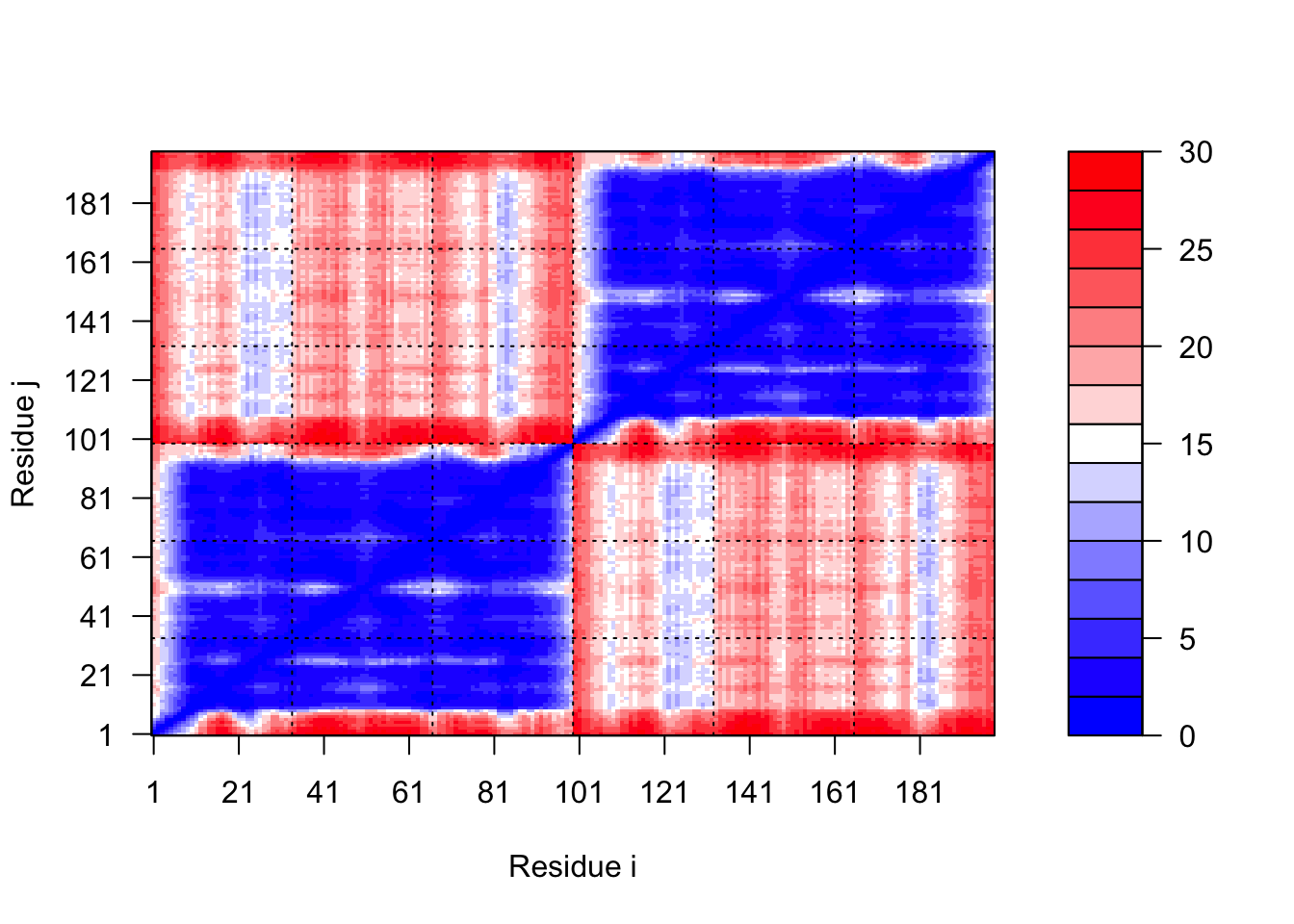
.a3m file:
aln_file <- list.files(path = results_dir,
pattern = "\\.a3m$",
full.names = TRUE)
aln <- read.fasta(aln_file[1], to.upper = TRUE)[1] " ** Duplicated sequence id's: 101 **"
[2] " ** Duplicated sequence id's: 101 **"dim(aln$ali)[1] 5397 132sim <- conserv(aln)
plotb3(sim[1:99], ylab="Conservation Score", xlab="Residue Position")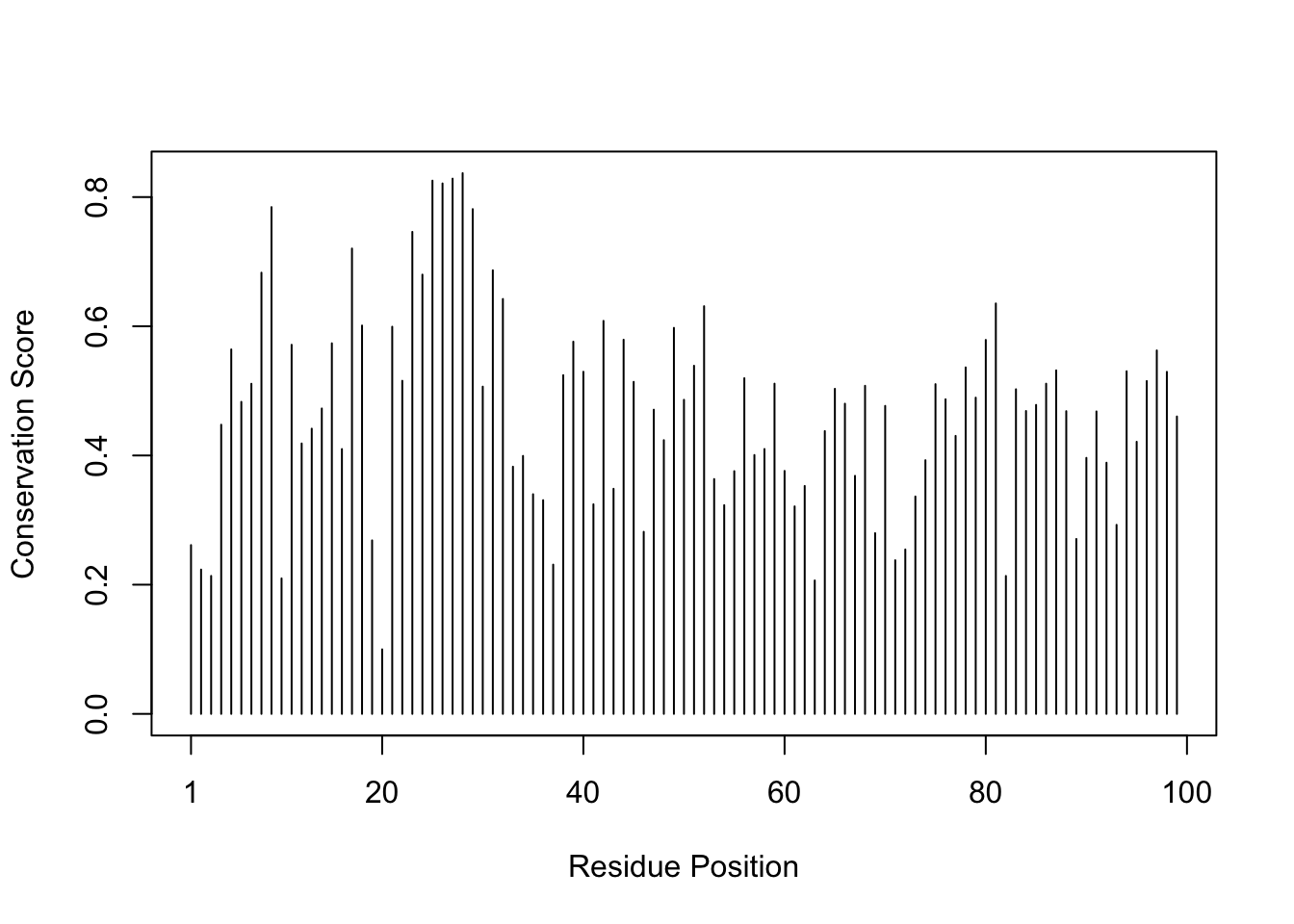
Core fit + RMSF
core <- core.find(pdbs) core size 197 of 198 vol = 9885.419
core size 196 of 198 vol = 6898.241
core size 195 of 198 vol = 1338.035
core size 194 of 198 vol = 1040.677
core size 193 of 198 vol = 951.865
core size 192 of 198 vol = 899.087
core size 191 of 198 vol = 834.733
core size 190 of 198 vol = 771.342
core size 189 of 198 vol = 733.069
core size 188 of 198 vol = 697.285
core size 187 of 198 vol = 659.748
core size 186 of 198 vol = 625.28
core size 185 of 198 vol = 589.548
core size 184 of 198 vol = 568.261
core size 183 of 198 vol = 545.022
core size 182 of 198 vol = 512.897
core size 181 of 198 vol = 490.731
core size 180 of 198 vol = 470.274
core size 179 of 198 vol = 450.738
core size 178 of 198 vol = 434.743
core size 177 of 198 vol = 420.345
core size 176 of 198 vol = 406.666
core size 175 of 198 vol = 393.341
core size 174 of 198 vol = 382.402
core size 173 of 198 vol = 372.866
core size 172 of 198 vol = 357.001
core size 171 of 198 vol = 346.576
core size 170 of 198 vol = 337.454
core size 169 of 198 vol = 326.668
core size 168 of 198 vol = 314.959
core size 167 of 198 vol = 304.136
core size 166 of 198 vol = 294.561
core size 165 of 198 vol = 285.658
core size 164 of 198 vol = 278.893
core size 163 of 198 vol = 266.773
core size 162 of 198 vol = 259.003
core size 161 of 198 vol = 247.731
core size 160 of 198 vol = 239.849
core size 159 of 198 vol = 234.973
core size 158 of 198 vol = 230.071
core size 157 of 198 vol = 221.995
core size 156 of 198 vol = 215.629
core size 155 of 198 vol = 206.8
core size 154 of 198 vol = 196.992
core size 153 of 198 vol = 188.547
core size 152 of 198 vol = 182.27
core size 151 of 198 vol = 176.961
core size 150 of 198 vol = 170.72
core size 149 of 198 vol = 166.128
core size 148 of 198 vol = 159.805
core size 147 of 198 vol = 153.775
core size 146 of 198 vol = 149.101
core size 145 of 198 vol = 143.664
core size 144 of 198 vol = 137.145
core size 143 of 198 vol = 132.523
core size 142 of 198 vol = 127.237
core size 141 of 198 vol = 121.579
core size 140 of 198 vol = 116.78
core size 139 of 198 vol = 112.575
core size 138 of 198 vol = 108.175
core size 137 of 198 vol = 105.137
core size 136 of 198 vol = 101.254
core size 135 of 198 vol = 97.379
core size 134 of 198 vol = 92.978
core size 133 of 198 vol = 88.188
core size 132 of 198 vol = 84.032
core size 131 of 198 vol = 81.902
core size 130 of 198 vol = 78.023
core size 129 of 198 vol = 75.276
core size 128 of 198 vol = 73.057
core size 127 of 198 vol = 70.699
core size 126 of 198 vol = 68.976
core size 125 of 198 vol = 66.707
core size 124 of 198 vol = 64.376
core size 123 of 198 vol = 61.145
core size 122 of 198 vol = 59.029
core size 121 of 198 vol = 56.625
core size 120 of 198 vol = 54.369
core size 119 of 198 vol = 51.826
core size 118 of 198 vol = 49.651
core size 117 of 198 vol = 48.19
core size 116 of 198 vol = 46.644
core size 115 of 198 vol = 44.748
core size 114 of 198 vol = 43.288
core size 113 of 198 vol = 41.089
core size 112 of 198 vol = 39.143
core size 111 of 198 vol = 36.468
core size 110 of 198 vol = 34.114
core size 109 of 198 vol = 31.467
core size 108 of 198 vol = 29.445
core size 107 of 198 vol = 27.323
core size 106 of 198 vol = 25.82
core size 105 of 198 vol = 24.149
core size 104 of 198 vol = 22.647
core size 103 of 198 vol = 21.068
core size 102 of 198 vol = 19.953
core size 101 of 198 vol = 18.3
core size 100 of 198 vol = 15.723
core size 99 of 198 vol = 14.841
core size 98 of 198 vol = 11.646
core size 97 of 198 vol = 9.435
core size 96 of 198 vol = 7.354
core size 95 of 198 vol = 6.181
core size 94 of 198 vol = 5.667
core size 93 of 198 vol = 4.706
core size 92 of 198 vol = 3.664
core size 91 of 198 vol = 2.77
core size 90 of 198 vol = 2.151
core size 89 of 198 vol = 1.715
core size 88 of 198 vol = 1.15
core size 87 of 198 vol = 0.874
core size 86 of 198 vol = 0.685
core size 85 of 198 vol = 0.528
core size 84 of 198 vol = 0.37
FINISHED: Min vol ( 0.5 ) reachedcore.inds <- print(core, vol = 0.5)# 85 positions (cumulative volume <= 0.5 Angstrom^3)
start end length
1 9 49 41
2 52 95 44xyz <- pdbfit(pdbs, core.inds, outpath = "corefit_structures")
rf <- rmsf(xyz)
plotb3(rf, sse = pdb_ref, ylab="RMSF (Å)", xlab="Residue Position")
abline(v = 99, col = "gray")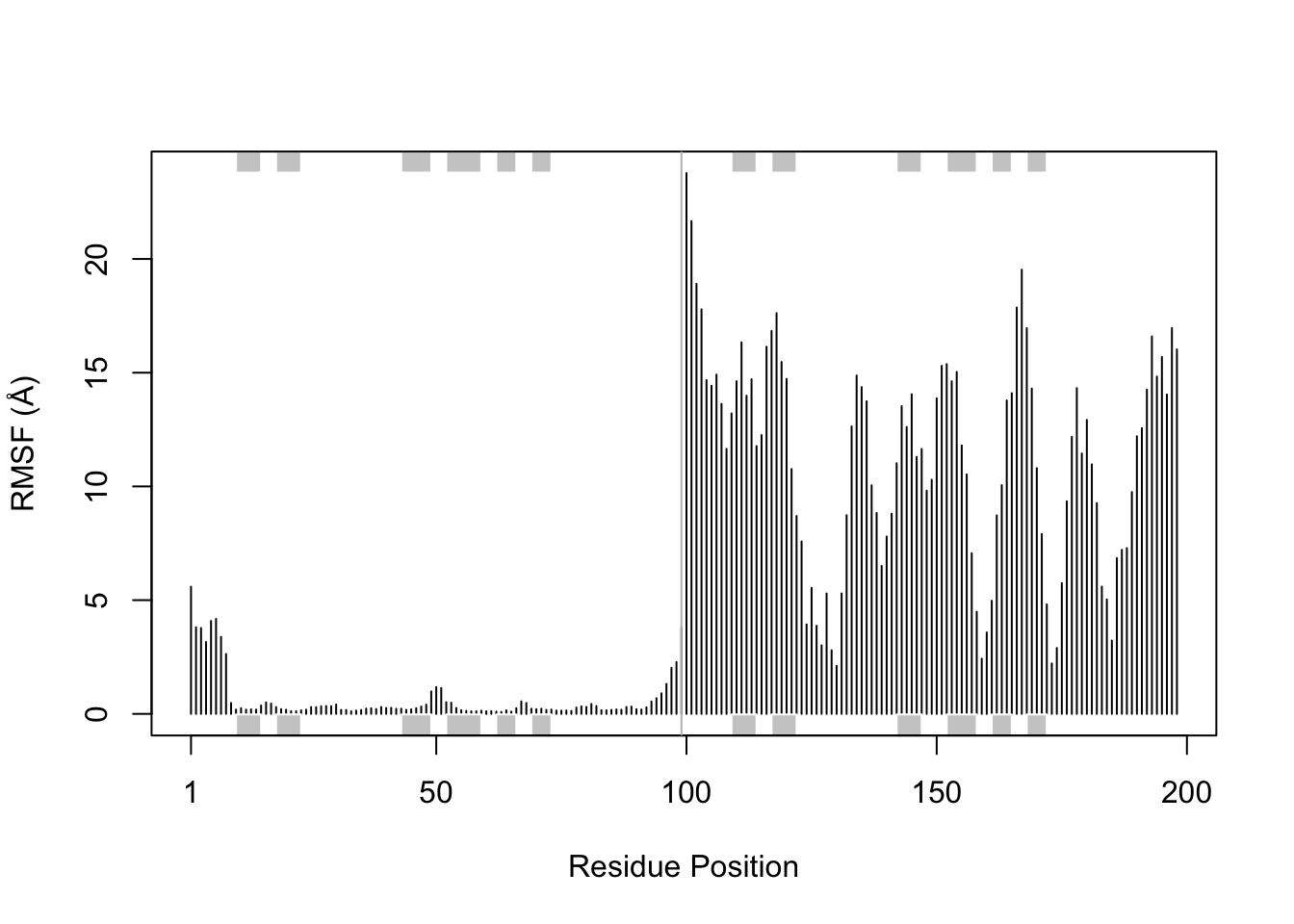
Mapping conservation onto a PDB for Mol
m1.pdb <- read.pdb(pdb_files[1])
occ <- vec2resno(c(sim[1:99], sim[1:99]), m1.pdb$atom$resno)
write.pdb(m1.pdb, o = occ, file = "m1_conserv.pdb")